Benchtop IR Imaging of Live Cells: Monitoring the Total Mass of Biomolecules in Single Cells
- PMID: 39230511
- PMCID: PMC11431153
- DOI: 10.1021/acs.analchem.4c02108
Benchtop IR Imaging of Live Cells: Monitoring the Total Mass of Biomolecules in Single Cells
Abstract
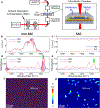
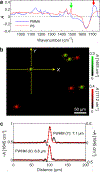
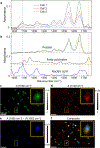
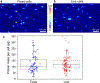
Absolute quantity imaging of biomolecules on a single cell level is critical for measurement assurance in biosciences and bioindustries. While infrared (IR) transmission microscopy is a powerful label-free imaging modality capable of chemical quantification, its applicability to hydrated biological samples remains challenging due to the strong IR absorption by water. Traditional IR imaging of hydrated cells relies on powerful light sources, such as synchrotrons, to mitigate the light absorption by water. However, we overcome this challenge by applying a solvent absorption compensation (SAC) technique to a home-built benchtop IR microscope based on an external-cavity quantum cascade laser. SAC-IR microscopy adjusts the incident light using a pair of polarizers to precompensate the IR absorption by water while retaining the full dynamic range. Integrating the IR absorbance over a cell yields the total mass of biomolecules per cell. We monitor the total mass of the biomolecules of live fibroblast cells over 12 h, demonstrating promise for advancing our understanding of the biomolecular processes occurring in live cells on the single-cell level.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

References
-
- Poulhazan A; Dickwella Widanage MC; Muszyński A; Arnold AA; Warschawski DE; Azadi P; Marcotte I; Wang T Identification and Quantification of Glycans in Whole Cells: Architecture of Microalgal Polysaccharides Described by Solid-State Nuclear Magnetic Resonance. J Am Chem Soc 2021, 143 (46), 19374–19388. 10.1021/JACS.1C07429. - DOI - PMC - PubMed
-
- Vallabh SM; Nobuhara CK; Llorens F; Zerr I; Parchi P; Capellari S; Kuhn E; Klickstein J; Safar JG; Nery FC; Swoboda KJ; Geschwind MD; Zetterberg H; Arnold SE; Minikel EV; Schreiber SL Prion Protein Quantification in Human Cerebrospinal Fluid as a Tool for Prion Disease Drug Development. Proc Natl Acad Sci U S A 2019, 116 (16), 7793–7798. 10.1073/PNAS.1901947116. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

