Metagenomic functional profiling: to sketch or not to sketch?
- PMID: 39230701
- PMCID: PMC11373326
- DOI: 10.1093/bioinformatics/btae397
Metagenomic functional profiling: to sketch or not to sketch?
Abstract
Motivation: Functional profiling of metagenomic samples is essential to decipher the functional capabilities of microbial communities. Traditional and more widely used functional profilers in the context of metagenomics rely on aligning reads against a known reference database. However, aligning sequencing reads against a large and fast-growing database is computationally expensive. In general, k-mer-based sketching techniques have been successfully used in metagenomics to address this bottleneck, notably in taxonomic profiling. In this work, we describe leveraging FracMinHash (implemented in sourmash, a publicly available software), a k-mer-sketching algorithm, to obtain functional profiles of metagenome samples.
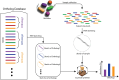
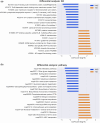
Results: We show how pieces of the sourmash software (and the resulting FracMinHash sketches) can be put together in a pipeline to functionally profile a metagenomic sample. We named our pipeline fmh-funprofiler. We report that the functional profiles obtained using this pipeline demonstrate comparable completeness and better purity compared to the profiles obtained using other alignment-based methods when applied to simulated metagenomic data. We also report that fmh-funprofiler is 39-99× faster in wall-clock time, and consumes up to 40-55× less memory. Coupled with the KEGG database, this method not only replicates fundamental biological insights but also highlights novel signals from the Human Microbiome Project datasets.
Availability and implementation: This fast and lightweight metagenomic functional profiler is freely available and can be accessed here: https://github.com/KoslickiLab/fmh-funprofiler. All scripts of the analyses we present in this manuscript can be found on GitHub.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures


References
-
- Broder AZ. On the resemblance and containment of documents. In: Proceedings of Compression and Complexity of SEQUENCES, Salerno, Italy, 1997 (Cat. No. 97TB100171). IEEE, 1997, 21–9.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

