Enumerative Discovery of Noncanonical Polypeptide Secondary Structures
- PMID: 39231524
- PMCID: PMC11421003
- DOI: 10.1021/jacs.4c04991
Enumerative Discovery of Noncanonical Polypeptide Secondary Structures
Abstract
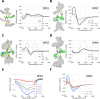
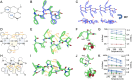
Energetically favorable local interactions can overcome the entropic cost of chain ordering and cause otherwise flexible polymers to adopt regularly repeating backbone conformations. A prominent example is the α helix present in many protein structures, which is stabilized by i, i + 4 hydrogen bonds between backbone peptide units. With the increased chemical diversity offered by unnatural amino acids and backbones, it has been possible to identify regularly repeating structures not present in proteins, but to date, there has been no systematic approach for identifying new polymers likely to have such structures despite their considerable potential for molecular engineering. Here we describe a systematic approach to search through dipeptide combinations of 130 chemically diverse amino acids to identify those predicted to populate unique low-energy states. We characterize ten newly identified dipeptide repeating structures using circular dichroism spectroscopy and comparison with calculated spectra. NMR and X-ray crystallographic structures of two of these dipeptide-repeat polymers are similar to the computational models. Our approach is readily generalizable to identify low-energy repeating structures for a wide variety of polymers, and our ordered dipeptide repeats provide new building blocks for molecular engineering.
Conflict of interest statement
The authors declare the following competing financial interest(s): GTM is a founder of Nexomics Biosciences, Inc, and APM, PJS, and DB are founders of Vilya, Inc. These do not represent conflicts of interest for this study.
Figures


References
-
- Gellman S. H. Foldamers: A Manifesto. Acc. Chem. Res. 1998, 31, 173–180. 10.1021/ar960298r. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

