Improving the Annotation Process in Computational Pathology: A Pilot Study with Manual and Semi-automated Approaches on Consumer and Medical Grade Devices
- PMID: 39231887
- PMCID: PMC11950598
- DOI: 10.1007/s10278-024-01248-x
Improving the Annotation Process in Computational Pathology: A Pilot Study with Manual and Semi-automated Approaches on Consumer and Medical Grade Devices
Abstract
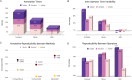
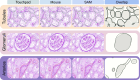
The development of reliable artificial intelligence (AI) algorithms in pathology often depends on ground truth provided by annotation of whole slide images (WSI), a time-consuming and operator-dependent process. A comparative analysis of different annotation approaches is performed to streamline this process. Two pathologists annotated renal tissue using semi-automated (Segment Anything Model, SAM)) and manual devices (touchpad vs mouse). A comparison was conducted in terms of working time, reproducibility (overlap fraction), and precision (0 to 10 accuracy rated by two expert nephropathologists) among different methods and operators. The impact of different displays on mouse performance was evaluated. Annotations focused on three tissue compartments: tubules (57 annotations), glomeruli (53 annotations), and arteries (58 annotations). The semi-automatic approach was the fastest and had the least inter-observer variability, averaging 13.6 ± 0.2 min with a difference (Δ) of 2%, followed by the mouse (29.9 ± 10.2, Δ = 24%), and the touchpad (47.5 ± 19.6 min, Δ = 45%). The highest reproducibility in tubules and glomeruli was achieved with SAM (overlap values of 1 and 0.99 compared to 0.97 for the mouse and 0.94 and 0.93 for the touchpad), though SAM had lower reproducibility in arteries (overlap value of 0.89 compared to 0.94 for both the mouse and touchpad). No precision differences were observed between operators (p = 0.59). Using non-medical monitors increased annotation times by 6.1%. The future employment of semi-automated and AI-assisted approaches can significantly speed up the annotation process, improving the ground truth for AI tool development.
Keywords: Annotation; Artificial intelligence; Computational pathology; Digital pathology; Segment Anything Model.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethical Approval: Approval was obtained from the local ethics committee (PNRR-MR1-2022–12375735, 03/16/23). Conflict of Interest: The authors declare no competing interests.
Figures


Similar articles
-
Overcoming an Annotation Hurdle: Digitizing Pen Annotations from Whole Slide Images.J Pathol Inform. 2021 Feb 23;12:9. doi: 10.4103/jpi.jpi_85_20. eCollection 2021. J Pathol Inform. 2021. PMID: 34012713 Free PMC article.
-
Annotation Practices in Computational Pathology: A European Society of Digital and Integrative Pathology (ESDIP) Survey Study.Lab Invest. 2025 Mar;105(3):102203. doi: 10.1016/j.labinv.2024.102203. Epub 2024 Nov 29. Lab Invest. 2025. PMID: 39615882
-
Semantic annotation for computational pathology: multidisciplinary experience and best practice recommendations.J Pathol Clin Res. 2022 Mar;8(2):116-128. doi: 10.1002/cjp2.256. Epub 2022 Jan 10. J Pathol Clin Res. 2022. PMID: 35014198 Free PMC article.
-
Artificial intelligence and algorithmic computational pathology: an introduction with renal allograft examples.Histopathology. 2021 May;78(6):791-804. doi: 10.1111/his.14304. Epub 2021 Mar 8. Histopathology. 2021. PMID: 33211332 Free PMC article. Review.
-
The potential of artificial intelligence-based applications in kidney pathology.Curr Opin Nephrol Hypertens. 2022 May 1;31(3):251-257. doi: 10.1097/MNH.0000000000000784. Epub 2022 Feb 14. Curr Opin Nephrol Hypertens. 2022. PMID: 35165248 Free PMC article. Review.
Cited by
-
Automating liver biopsy segmentation with a robust, open-source tool for pathology research: the HOTSPoT model.NPJ Digit Med. 2025 Jul 18;8(1):455. doi: 10.1038/s41746-025-01870-1. NPJ Digit Med. 2025. PMID: 40681745 Free PMC article.
-
Analyzing the performance of biomedical time-series segmentation with electrophysiology data.Sci Rep. 2025 Apr 6;15(1):11776. doi: 10.1038/s41598-025-90533-y. Sci Rep. 2025. PMID: 40189617 Free PMC article.
References
-
- Hanna, M. G., Ardon, O., Reuter, V. E., Sirintrapun, S. J., England, C., Klimstra, D. S. & Hameed, M. R. Integrating digital pathology into clinical practice. Mod. Pathol.35, 152–164 (2022). - PubMed
-
- Pisapia, P., L’Imperio, V., Galuppini, F., Sajjadi, E., Russo, A., Cerbelli, B., Fraggetta, F., d’Amati, G., Troncone, G., Fassan, M., Fusco, N., Pagni, F. & Malapelle, U. The evolving landscape of anatomic pathology. Crit. Rev. Oncol. Hematol.178, 103776 (2022). - PubMed
-
- L’Imperio, V., Casati, G., Cazzaniga, G., Tarabini, A., Bolognesi, M. M., Gibilisco, F., Fraggetta, F. & Pagni, F. Improvements in digital pathology equipment for renal biopsies: updating the standard model. J. Nephrol. (2023). 10.1007/s40620-023-01568-1 - PubMed
-
- Cazzaniga, G., Rossi, M., Eccher, A., Girolami, I., L’Imperio, V., Van Nguyen, H., Becker, J. U., Bueno García, M. G., Sbaraglia, M., Dei Tos, A. P., Gambaro, G. & Pagni, F. Time for a full digital approach in nephropathology: a systematic review of current artificial intelligence applications and future directions. J. Nephrol. (2023). 10.1007/s40620-023-01775-w - PMC - PubMed
MeSH terms
Grants and funding
- DIPLOMAT - PNRR-MR1-2022-12375735/Next Generation EU - NRRP M6C2
- Dipartimenti di Eccellenza 2023-2027 (l. 232/2016/Ministero dell'Istruzione, dell'Università e della Ricerca
- art. 1/Ministero dell'Istruzione, dell'Università e della Ricerca
- commi 314 - 337)/Ministero dell'Istruzione, dell'Università e della Ricerca
- GR-2021-12374235/Ministero dell'Università e della Ricerca
LinkOut - more resources
Full Text Sources
