Diverse viral cas genes antagonize CRISPR immunity
- PMID: 39232173
- PMCID: PMC11991930
- DOI: 10.1038/s41586-024-07923-x
Diverse viral cas genes antagonize CRISPR immunity
Abstract
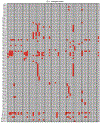
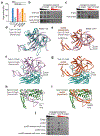
Prokaryotic CRISPR-Cas immunity is subverted by anti-CRISPRs (Acrs), which inhibit Cas protein activities when expressed during the phage lytic cycle or from resident prophages or plasmids1. Acrs often bind to specific cognate Cas proteins, and hence inhibition is typically limited to a single CRISPR-Cas subtype2. Furthermore, although acr genes are frequently organized together in phage-associated gene clusters3, how such inhibitors initially evolve has remained unclear. Here we investigated the Acr content and inhibition specificity of diverse Listeria isolates, which naturally harbour four CRISPR-Cas systems (types I-B, II-A, II-C and VI-A). We observed widespread antagonism of CRISPR, which we traced to 11 previously unknown and 4 known acr gene families encoded by endogenous mobile elements. Among these were two Acrs that possess sequence homology to type I-B Cas proteins, one of which assembles into a defective interference complex. Surprisingly, an additional type I-B Cas homologue did not affect type I immunity, but instead inhibited the RNA-targeting type VI CRISPR system by means of CRISPR RNA (crRNA) degradation. By probing viral sequence databases, we detected abundant orphan cas genes located within putative anti-defence gene clusters. Among them, we verified the activity of a particularly broad-spectrum cas3 homologue that inhibits type I-B, II-A and VI-A CRISPR immunity. Our observations provide direct evidence of Acr evolution by cas gene co-option, and new genes with potential for broad-spectrum control of genome editing technologies.
© 2024. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
A.J.M. is a co-founder of Profluent Bio. J.B.-D. is a scientific advisory board member of SNIPR Biome and Excision Biotherapeutics, a consultant to LeapFrog Bio and a scientific advisory board member and co-founder of Acrigen Biosciences. The Bondy-Denomy lab received research support from Felix Biotechnology. The other authors declare no competing interests.
Figures















References
-
- Dong L. et al. An anti-CRISPR protein disables type V Cas12a by acetylation. Nat. Struct. Mol. Biol 26, 308–314 (2019). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

