Disorder-mediated interactions target proteins to specific condensates
- PMID: 39232584
- PMCID: PMC11416317
- DOI: 10.1016/j.molcel.2024.08.017
Disorder-mediated interactions target proteins to specific condensates
Abstract
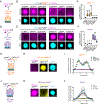
Selective compartmentalization of cellular contents is fundamental to the regulation of biochemistry. Although membrane-bound organelles control composition by using a semi-permeable barrier, biomolecular condensates rely on interactions among constituents to determine composition. Condensates are formed by dynamic multivalent interactions, often involving intrinsically disordered regions (IDRs) of proteins, yet whether distinct compositions can arise from these dynamic interactions is not known. Here, by comparative analysis of proteins differentially partitioned by two different condensates, we find that distinct compositions arise through specific IDR-mediated interactions. The IDRs of differentially partitioned proteins are necessary and sufficient for selective partitioning. Distinct sequence features are required for IDRs to partition, and swapping these sequence features changes the specificity of partitioning. Swapping whole IDRs retargets proteins and their biochemical activity to different condensates. Our results demonstrate that IDR-mediated interactions can target proteins to specific condensates, enabling the spatial regulation of biochemistry within the cell.
Keywords: IDR; biomolecular condensates; condensate composition; condensate function; intrinsically disordered regions; nuclear organization; specificity.
Copyright © 2024 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests B.R.S. is an advisory board member of Molecular Cell. B.A.C. and M.O. were employed by Pfizer Inc. at the time of writing.
Figures




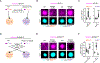

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

