LIG1 is a novel marker for bladder cancer prognosis: evidence based on experimental studies, machine learning and single-cell sequencing
- PMID: 39234248
- PMCID: PMC11371609
- DOI: 10.3389/fimmu.2024.1419126
LIG1 is a novel marker for bladder cancer prognosis: evidence based on experimental studies, machine learning and single-cell sequencing
Abstract
Background: Bladder cancer, a highly fatal disease, poses a significant threat to patients. Positioned at 19q13.2-13.3, LIG1, one of the four DNA ligases in mammalian cells, is frequently deleted in tumour cells of diverse origins. Despite this, the precise involvement of LIG1 in BLCA remains elusive. This pioneering investigation delves into the uncharted territory of LIG1's impact on BLCA. Our primary objective is to elucidate the intricate interplay between LIG1 and BLCA, alongside exploring its correlation with various clinicopathological factors.
Methods: We retrieved gene expression data of para-carcinoma tissues and bladder cancer (BLCA) from the GEO repository. Single-cell sequencing data were processed using the "Seurat" package. Differential expression analysis was then performed with the "Limma" package. The construction of scale-free gene co-expression networks was achieved using the "WGCNA" package. Subsequently, a Venn diagram was utilized to extract genes from the positively correlated modules identified by WGCNA and intersect them with differentially expressed genes (DEGs), isolating the overlapping genes. The "STRINGdb" package was employed to establish the protein-protein interaction (PPI) network.Hub genes were identified through the PPI network using the Betweenness Centrality (BC) algorithm. We conducted KEGG and GO enrichment analyses to uncover the regulatory mechanisms and biological functions associated with the hub genes. A machine-learning diagnostic model was established using the R package "mlr3verse." Mutation profiles between the LIG1^high and LIG1^low groups were visualized using the BEST website. Survival analyses within the LIG1^high and LIG1^low groups were performed using the BEST website and the GENT2 website. Finally, a series of functional experiments were executed to validate the functional role of LIG1 in BLCA.
Results: Our investigation revealed an upregulation of LIG1 in BLCA specimens, with heightened LIG1 levels correlating with unfavorable overall survival outcomes. Functional enrichment analysis of hub genes, as evidenced by GO and KEGG enrichment analyses, highlighted LIG1's involvement in critical function such as the DNA replication, cellular senescence, cell cycle and the p53 signalling pathway. Notably, the mutational landscape of BLCA varied significantly between LIG1high and LIG1low groups.Immune infiltrating analyses suggested a pivotal role for LIG1 in immune cell recruitment and immune regulation within the BLCA microenvironment, thereby impacting prognosis. Subsequent experimental validations further underscored the significance of LIG1 in BLCA pathogenesis, consolidating its functional relevance in BLCA samples.
Conclusions: Our research demonstrates that LIG1 plays a crucial role in promoting bladder cancer malignant progression by heightening proliferation, invasion, EMT, and other key functions, thereby serving as a potential risk biomarker.
Keywords: LIG1; bioinformatics; machine-learning; single-cell; tumorinfiltrating immune cell; urothelial bladder cancer.
Copyright © 2024 Song, Shen, Feng, He, Chen, Zhang, Luo, Han, Tong and Jin.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures







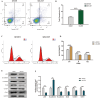
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

