ATRX restricts Human Cytomegalovirus (HCMV) viral DNA replication through heterochromatinization and minimizes unpackaged viral genomes
- PMID: 39236084
- PMCID: PMC11407672
- DOI: 10.1371/journal.ppat.1012516
ATRX restricts Human Cytomegalovirus (HCMV) viral DNA replication through heterochromatinization and minimizes unpackaged viral genomes
Abstract
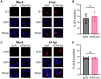
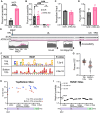
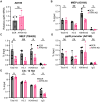
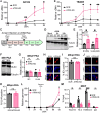
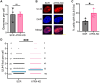
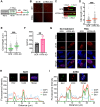
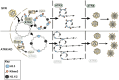
ATRX limits the accumulation of human cytomegalovirus (HCMV) Immediate Early (IE) proteins at the start of productive, lytic infections, and thus is a part of the cell-intrinsic defenses against infecting viruses. ATRX is a chromatin remodeler and a component of a histone chaperone complex. Therefore, we hypothesized ATRX would inhibit the transcription of HCMV IE genes by increasing viral genome heterochromatinization and decreasing its accessibility. To test this hypothesis, we quantitated viral transcription and genome structure in cells replete with or depleted of ATRX. We found ATRX did indeed limit viral IE transcription, increase viral genome chromatinization, and decrease viral genome accessibility. The inhibitory effects of ATRX extended to Early (E) and Late (L) viral protein accumulation, viral DNA replication, and progeny virion output. However, we found the negative effects of ATRX on HCMV viral DNA replication were independent of its effects on viral IE and E protein accumulation but correlated with viral genome heterochromatinization. Interestingly, the increased number of viral genomes synthesized in ATRX-depleted cells were not efficiently packaged, indicating the ATRX-mediated restriction to HCMV viral DNA replication may benefit productive infection by increasing viral fitness. Our work mechanistically describes the antiviral function of ATRX and introduces a novel, pro-viral role for this protein, perhaps explaining why, unlike during infections with other herpesviruses, it is not directly targeted by a viral countermeasure in HCMV infected cells.
Copyright: © 2024 Walter et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

