YIPF2 regulates genome integrity
- PMID: 39238039
- PMCID: PMC11376028
- DOI: 10.1186/s13578-024-01300-x
YIPF2 regulates genome integrity
Abstract
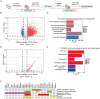
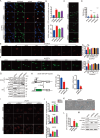
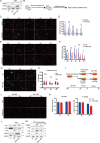
Understanding of the mechanisms for genome integrity maintenance can help in developing effective intervention strategies to combat aging. A whole-genome RNAi screen was conducted to identify novel factors involved in maintaining genome stability. The potential target genes identified in the screening are related to the cell cycle, proteasome, and spliceosomes. Unexpectedly, the Golgi protein YIPF2 has been found to play a critical role in maintaining genome stability. The depletion of YIPF2 hinders the process of homologous recombination (HR) repair, which then triggers DNA damage response mechanisms, ultimately leading to cellular senescence. The overexpression of YIPF2 facilitated cellular recovery from DNA damage induced by chemotherapy agents or replicative senescence-associated DNA damage. Our findings indicate that only the intact Golgi apparatus containing YIPF2 provides a protective effect on genome integrity.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

References
Grants and funding
LinkOut - more resources
Full Text Sources

