Prenatal opioid exposure significantly impacts placental protein kinase C (PKC) and drug transporters, leading to drug resistance and neonatal opioid withdrawal syndrome
- PMID: 39238930
- PMCID: PMC11376091
- DOI: 10.3389/fnins.2024.1442915
Prenatal opioid exposure significantly impacts placental protein kinase C (PKC) and drug transporters, leading to drug resistance and neonatal opioid withdrawal syndrome
Abstract
Background: Neonatal Opioid Withdrawal Syndrome (NOWS) is a consequence of in-utero exposure to prenatal maternal opioids, resulting in the manifestation of symptoms like irritability, feeding problems, tremors, and withdrawal signs. Opioid use disorder (OUD) during pregnancy can profoundly impact both mother and fetus, disrupting fetal brain neurotransmission and potentially leading to long-term neurological, behavioral, and vision issues, and increased infant mortality. Drug resistance complicates OUD and NOWS treatment, with protein kinase regulation of drug transporters not fully understood.
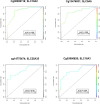
Methods: DNA methylation levels of ATP-binding cassette (ABC) and solute carrier (SLC) drug transporters, along with protein kinase C (PKC) genes, were assessed in 96 placental samples using the Illumina Infinium MethylationEPIC array (850K). Samples were collected from three distinct groups: 32 mothers with infants prenatally exposed to opioids who needed pharmacological intervention for NOWS, 32 mothers with prenatally opioid-exposed infants who did not necessitate NOWS treatment, and 32 mothers who were not exposed to opioids during pregnancy.
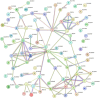
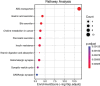
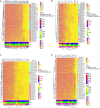
Results: We identified 69 significantly differentially methylated SLCs, with 24 hypermethylated and 34 hypomethylated, and 11 exhibiting both types of methylation changes including SLC13A3, SLC15A2, SLC16A11, SLC16A3, SLC19A2, and SLC26A1. We identified methylation changes in 11 ABC drug transporters (ABCA1, ABCA12, ABCA2, ABCB10, ABCB5, ABCC12, ABCC2, ABCC9, ABCE1, ABCC7, ABCB3): 3 showed hypermethylation, 3 hypomethylation, and 5 exhibited both. Additionally, 7 PKC family genes (PRKCQ, PRKAA1, PRKCA, PRKCB, PRKCH, PRKCI, and PRKCZ) showed methylation changes. These genes are associated with 13 pathways involved in NOWS, including ABC transporters, bile secretion, pancreatic secretion, insulin resistance, glutamatergic synapse, and gastric acid secretion.
Conclusion: We report epigenetic changes in PKC-related regulation of drug transporters, which could improve our understanding of clinical outcomes like drug resistance, pharmacokinetics, drug-drug interactions, and drug toxicity, leading to maternal relapse and severe NOWS. Novel drugs targeting PKC pathways and transporters may improve treatment outcomes for OUD in pregnancy and NOWS.
Keywords: bile secretion; biomarker; drug transporters; insulin resistance; neonatal opioid withdrawal syndrome SLC transporters ABC transporters; opioid use; pancreatic secretion; protein kinases C.
Copyright © 2024 Radhakrishna, Radhakrishnan, Uppala, Muvvala, Prajapati, Rawal, Bahado-Singh and Sadhasivam.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures

Similar articles
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Mothers with Concurrent Opioid and Cocaine Use and Neonatal Opioid Withdrawal Syndrome.Children (Basel). 2025 Jul 11;12(7):916. doi: 10.3390/children12070916. Children (Basel). 2025. PMID: 40723109 Free PMC article.
-
Adapting a Risk Prediction Tool for Neonatal Opioid Withdrawal Syndrome.Pediatrics. 2025 Apr 1;155(4):e2024068673. doi: 10.1542/peds.2024-068673. Pediatrics. 2025. PMID: 40024274
-
Acupuncture for neonatal abstinence syndrome in newborn infants.Cochrane Database Syst Rev. 2025 Feb 21;2(2):CD014160. doi: 10.1002/14651858.CD014160.pub2. Cochrane Database Syst Rev. 2025. PMID: 39981752
-
Prenatal interventions for congenital diaphragmatic hernia for improving outcomes.Cochrane Database Syst Rev. 2015 Nov 27;2015(11):CD008925. doi: 10.1002/14651858.CD008925.pub2. Cochrane Database Syst Rev. 2015. PMID: 26611822 Free PMC article.
Cited by
-
Prenatal opioid exposure alters pain perception and increases long-term health risks in infants with neonatal opioid withdrawal syndrome.Front Pain Res (Lausanne). 2025 Apr 17;6:1497801. doi: 10.3389/fpain.2025.1497801. eCollection 2025. Front Pain Res (Lausanne). 2025. PMID: 40313396 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

