Early moderate prenatal alcohol exposure and maternal diet impact offspring DNA methylation across species
- PMID: 39239947
- PMCID: PMC11379454
- DOI: 10.7554/eLife.92135
Early moderate prenatal alcohol exposure and maternal diet impact offspring DNA methylation across species
Abstract
Alcohol consumption in pregnancy can affect genome regulation in the developing offspring but results have been contradictory. We employed a physiologically relevant murine model of short-term moderate prenatal alcohol exposure (PAE) resembling common patterns of alcohol consumption in pregnancy in humans. Early moderate PAE was sufficient to affect site-specific DNA methylation in newborn pups without altering behavioural outcomes in adult littermates. Whole-genome bisulfite sequencing of neonatal brain and liver revealed stochastic influence on DNA methylation that was mostly tissue-specific, with some perturbations likely originating as early as gastrulation. DNA methylation differences were enriched in non-coding genomic regions with regulatory potential indicative of broad effects of alcohol on genome regulation. Replication studies in human cohorts with fetal alcohol spectrum disorder suggested some effects were metastable at genes linked to disease-relevant traits including facial morphology, intelligence, educational attainment, autism, and schizophrenia. In our murine model, a maternal diet high in folate and choline protected against some of the damaging effects of early moderate PAE on DNA methylation. Our studies demonstrate that early moderate exposure is sufficient to affect fetal genome regulation even in the absence of overt phenotypic changes and highlight a role for preventative maternal dietary interventions.
Keywords: DNA methylation; alcohol; developmental biology; epigeneome; genetics; genomics; human; mouse.
Plain language summary
Drinking excessive amounts of alcohol during pregnancy can cause foetal alcohol spectrum disorder and other conditions in children that affect their physical and mental development. Many countries advise women who are pregnant or trying to conceive to avoid drinking alcohol entirely. However, surveys of large groups of women in Western countries indicate that most women continue drinking low to moderate amounts of alcohol until they discover they are pregnant and then stop consuming alcohol for the rest of their pregnancy. It remains unclear how this common drinking pattern affects the foetus. The instructions needed to build and maintain a human body are stored within molecules of DNA. Some regions of DNA called genes contain the instructions to make proteins, which perform many tasks in the body. Other so-called ‘non-coding’ regions do not code for any proteins but instead have roles in regulating gene activity. One way cells control which genes are switched on or off is adding or removing tags known as methyl groups to certain locations on DNA. Previous studies indicate that alcohol may affect how children develop by changing the patterns of methyl tags on DNA. To investigate the effect of moderate drinking during the early stages of pregnancy, Bestry et al. exposed pregnant mice to alcohol and examined how this affected the patterns of methyl tags on DNA in their offspring. The experiments found moderate levels of alcohol were sufficient to alter the patterns of methyl tags in the brains and livers of the newborn mice. Most of the changes were observed in non-coding regions of DNA, suggesting alcohol may affect how large groups of genes are regulated. Fewer changes in the patterns of methyl tags were found in mice whose mothers had diets rich in two essential nutrients known as folate and choline. Further experiments found that some of the affected mouse genes were similar to genes linked to foetal alcohol spectrum disorder and other related conditions in humans. These findings highlight the potential risks of consuming even moderate levels of alcohol during pregnancy and suggest that a maternal diet rich in folate and choline may help mitigate some of the harmful effects on the developing foetus.
© 2023, Bestry et al.
Conflict of interest statement
MB, AL, NK, EC, CB, JF, EE, JC, EM, JH, DH, SB, RL, MS, DM No competing interests declared
Figures

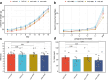

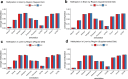
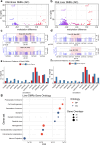
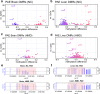

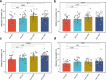
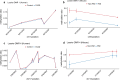
Update of
- doi: 10.1101/2023.06.15.544516
- doi: 10.7554/eLife.92135.1
- doi: 10.7554/eLife.92135.2
References
-
- Autism Spectrum Disorders Working Group of The Psychiatric Genomics Consortium Meta-analysis of GWAS of over 16,000 individuals with autism spectrum disorder highlights a novel locus at 10q24.32 and a significant overlap with schizophrenia. Molecular Autism. 2017;8:21. doi: 10.1186/s13229-017-0137-9. - DOI - PMC - PubMed
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

