Mother's milk microbiota is associated with the developing gut microbial consortia in very-low-birth-weight infants
- PMID: 39243753
- PMCID: PMC11525026
- DOI: 10.1016/j.xcrm.2024.101729
Mother's milk microbiota is associated with the developing gut microbial consortia in very-low-birth-weight infants
Abstract
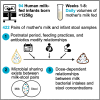
Mother's milk contains diverse bacterial communities, although their impact on microbial colonization in very-low-birth-weight (VLBW, <1,500 g) infants remains unknown. Here, we examine relationships between the microbiota in preterm mother's milk and the VLBW infant gut across initial hospitalization (n = 94 mother-infant dyads, 422 milk-stool pairs). Shared zero-radius operational taxonomic units (zOTUs) between milk-stool pairs account for ∼30%-40% of zOTUs in the VLBW infant's gut. We show dose-response relationships between intakes of several genera from milk and their concentrations in the infant's gut. These relationships and those related to microbial sharing change temporally and are modified by in-hospital feeding practices (especially direct breastfeeding) and maternal-infant antibiotic use. Correlations also exist between milk and stool microbial consortia, suggesting that multiple milk microbes may influence overall gut communities together. These results highlight that the mother's milk microbiota may shape the gut colonization of VLBW infants by delivering specific bacteria and through intricate microbial interactions.
Keywords: antibiotics; direct breastfeeding; donor milk; human milk; microbiome; microbiota; mother’s milk; nutrient fortification; preterm infant; very-low-birth-weight infant.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests E.M.C. acknowledges research support from Lallemand Health Solutions and Ocean Spray and consultant fees, speaker, and/or travel support from Danone, Nestlé, and Lallemand Health Solutions. P.M.S. is a stockholder and an advisory board member for Antibe Therapeutics Inc. A.S. is a co-founder of MedBiome, a clinical microbiomics company.
Figures







References
-
- Aguilar-lopez M., Dinsmoor A.M., Ho T.T.B., Donovan S.M., Dinsmoor A.M., Ho T.T.B., Sharon M., Aguilar-lopez M. A systematic review of the factors influencing microbial colonization of the preterm infant gut preterm infant gut. Gut Microb. 2021;13:1–33. doi: 10.1080/19490976.2021.1884514. - DOI - PMC - PubMed
-
- Pammi M., Cope J., Tarr P.I., Warner B.B., Morrow A.L., Mai V., Gregory K.E., Kroll J.S., McMurtry V., Ferris M.J., et al. Intestinal dysbiosis in preterm infants preceding necrotizing enterocolitis: A systematic review and meta-analysis. Microbiome. 2017;5 doi: 10.1186/s40168-017-0248-8. - DOI - PMC - PubMed
-
- Stewart C.J., Embleton N.D., Marrs E.C.L., Smith D.P., Fofanova T., Nelson A., Skeath T., Perry J.D., Petrosino J.F., Berrington J.E., Cummings S.P. Longitudinal development of the gut microbiome and metabolome in preterm neonates with late onset sepsis and healthy controls. Microbiome. 2017;5:75. doi: 10.1186/s40168-017-0295-1. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

