An open-source tool for automated human-level circling behavior detection
- PMID: 39245735
- PMCID: PMC11381541
- DOI: 10.1038/s41598-024-71665-z
An open-source tool for automated human-level circling behavior detection
Abstract
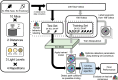
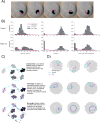
Quantitatively relating behavior to underlying biology is crucial in life science. Although progress in keypoint tracking tools has reduced barriers to recording postural data, identifying specific behaviors from this data remains challenging. Manual behavior coding is labor-intensive and inconsistent, while automatic methods struggle to explicitly define complex behaviors, even when they seem obvious to the human eye. Here, we demonstrate an effective technique for detecting circling in mice, a form of locomotion characterized by stereotyped spinning. Despite circling's extensive history as a behavioral marker, there currently exists no standard automated detection method. We developed a circling detection technique using simple postprocessing of keypoint data obtained from videos of freely-exploring (Cib2-/-;Cib3-/-) mutant mice, a strain previously found to exhibit circling behavior. Our technique achieves statistical parity with independent human observers in matching occurrence times based on human consensus, and it accurately distinguishes between videos of wild type mice and mutants. Our pipeline provides a convenient, noninvasive, quantitative tool for analyzing circling mouse models without the need for software engineering experience. Additionally, as the concepts underlying our approach are agnostic to the behavior being analyzed, and indeed to the modality of the recorded data, our results support the feasibility of algorithmically detecting specific research-relevant behaviors using readily-interpretable parameters tuned on the basis of human consensus.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Update of
-
An Open-Source Tool for Automated Human-Level Circling Behavior Detection.bioRxiv [Preprint]. 2023 May 30:2023.05.30.540066. doi: 10.1101/2023.05.30.540066. bioRxiv. 2023. Update in: Sci Rep. 2024 Sep 8;14(1):20914. doi: 10.1038/s41598-024-71665-z. PMID: 37398316 Free PMC article. Updated. Preprint.
References
-
- Mathis, A. et al. DeepLabCut: Markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci.21, 1281–1289 https://doi.org/10.1038/s41593-018-0209-y (2018). 10.1038/s41593-018-0209-y - DOI - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

