USP26 as a hepatitis B virus-induced deubiquitinase primes hepatocellular carcinogenesis by epigenetic remodeling
- PMID: 39251623
- PMCID: PMC11385750
- DOI: 10.1038/s41467-024-52201-z
USP26 as a hepatitis B virus-induced deubiquitinase primes hepatocellular carcinogenesis by epigenetic remodeling
Abstract
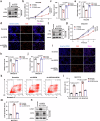
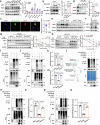
Despite recent advances in systemic therapy for hepatocellular carcinoma (HCC), the prognosis of hepatitis B virus (HBV)-induced HCC patients remains poor. By screening a sgRNA library targeting human deubiquitinases, we find that ubiquitin-specific peptidase 26 (USP26) deficiency impairs HBV-positive HCC cell proliferation. Genetically engineered murine models with Usp26 knockout confirm that Usp26 drives HCC tumorigenesis. Mechanistically, we find that the HBV-encoded protein HBx binds to the promoter and induces the production of USP26, which is an X-linked gene exclusively expressed in the testis. HBx consequently promotes the association of USP26 with SIRT1 to synergistically stabilize SIRT1 by deubiquitination, which promotes cell proliferation and impedes cell apoptosis to accelerate HCC tumorigenesis. In patients with HBV-positive HCC, USP26 is robustly induced, and its levels correlate with SIRT1 levels and poor prognosis. Collectively, our study highlights a causative link between HBV infection, deubiquitinase induction and development of HCC, identifying a druggable target, USP26.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

