Identification of cross-talk pathways and PANoptosis-related genes in periodontitis and Alzheimer's disease by bioinformatics analysis and machine learning
- PMID: 39258145
- PMCID: PMC11384588
- DOI: 10.3389/fnagi.2024.1430290
Identification of cross-talk pathways and PANoptosis-related genes in periodontitis and Alzheimer's disease by bioinformatics analysis and machine learning
Abstract
Background and objectives: Periodontitis (PD), a chronic inflammatory disease, is a serious threat to oral health and is one of the risk factors for Alzheimer's disease (AD). A growing body of evidence suggests that the two diseases are closely related. However, current studies have not provided a comprehensive understanding of the common genes and common mechanisms between PD and AD. This study aimed to screen the crosstalk genes of PD and AD and the potential relationship between cross-talk and PANoptosis-related genes. The relationship between core genes and immune cells will be analyzed to provide new targets for clinical treatment.
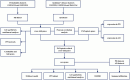
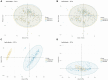
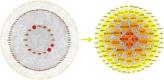
Materials and methods: The PD and AD datasets were downloaded from the GEO database and differential expression analysis was performed to obtain DEGs. Overlapping DEGs had cross-talk genes linking PD and OP, and PANoptosis-related genes were obtained from a literature review. Pearson coefficients were used to compute cross-talk and PANoptosis-related gene correlations in the PD and AD datasets. Cross-talk genes were obtained from the intersection of PD and AD-related genes, protein-protein interaction(PPI) networks were constructed and cross-talk genes were identified using the STRING database. The intersection of cross-talk and PANoptosis-related genes was defined as cross-talk-PANoptosis genes. Core genes were screened using ROC analysis and XGBoost. PPI subnetwork, gene-biological process, and gene-pathway networks were constructed based on the core genes. In addition, immune infiltration on the PD and AD datasets was analyzed using the CIBERSORT algorithm.
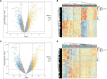
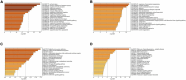
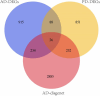
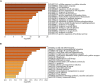
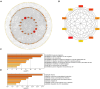
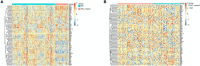
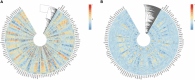
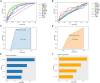
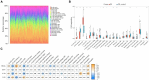
Results: 366 cross-talk genes were overlapping between PD DEGs and AD DEGs. The intersection of cross-talk genes with 109 PANoptosis-related genes was defined as cross-talk-PANoptosis genes. ROC and XGBoost showed that MLKL, DCN, IL1B, and IL18 were more accurate than the other cross-talk-PANoptosis genes in predicting the disease, as well as better in overall characterization. GO and KEGG analyses showed that the four core genes were involved in immunity and inflammation in the organism. Immune infiltration analysis showed that B cells naive, Plasma cells, and T cells gamma delta were significantly differentially expressed in patients with PD and AD compared with the normal group. Finally, 10 drugs associated with core genes were retrieved from the DGIDB database.
Conclusion: This study reveals the joint mechanism between PD and AD associated with PANoptosis. Analyzing the four core genes and immune cells may provide new therapeutic directions for the pathogenesis of PD combined with AD.
Keywords: Alzheimer’s disease; PANoptosis; common genes; immune infiltration; periodontitis.
Copyright © 2024 Chen, Dai, Li, Xin, Zou, Wang and Zhang and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Screening of crosstalk and pyroptosis-related genes linking periodontitis and osteoporosis based on bioinformatics and machine learning.Front Immunol. 2022 Aug 5;13:955441. doi: 10.3389/fimmu.2022.955441. eCollection 2022. Front Immunol. 2022. PMID: 35990678 Free PMC article.
-
Potential diagnostic markers and therapeutic targets for periodontitis and Alzheimer's disease based on bioinformatics analysis.J Periodontal Res. 2024 Apr;59(2):366-380. doi: 10.1111/jre.13220. Epub 2024 Jan 8. J Periodontal Res. 2024. PMID: 38189472
-
Transcriptomic analysis reveals molecular characterization and immune landscape of PANoptosis-related genes in atherosclerosis.Inflamm Res. 2024 Jun;73(6):961-978. doi: 10.1007/s00011-024-01877-6. Epub 2024 Apr 8. Inflamm Res. 2024. PMID: 38587531
-
Identification of Cross-Talk and Pyroptosis-Related Genes Linking Periodontitis and Rheumatoid Arthritis Revealed by Transcriptomic Analysis.Dis Markers. 2021 Dec 29;2021:5074305. doi: 10.1155/2021/5074305. eCollection 2021. Dis Markers. 2021. PMID: 35003389 Free PMC article.
-
Identification of early Alzheimer's disease subclass and signature genes based on PANoptosis genes.Front Immunol. 2024 Nov 22;15:1462003. doi: 10.3389/fimmu.2024.1462003. eCollection 2024. Front Immunol. 2024. PMID: 39650656 Free PMC article.
Cited by
-
Emerging regulated cell death mechanisms in bone remodeling: decoding ferroptosis, cuproptosis, disulfidptosis, and PANoptosis as therapeutic targets for skeletal disorders.Cell Death Discov. 2025 Jul 21;11(1):335. doi: 10.1038/s41420-025-02633-3. Cell Death Discov. 2025. PMID: 40691135 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

