Elasticity of the HIV-1 core facilitates nuclear entry and infection
- PMID: 39259747
- PMCID: PMC11419384
- DOI: 10.1371/journal.ppat.1012537
Elasticity of the HIV-1 core facilitates nuclear entry and infection
Abstract
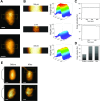
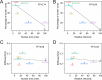
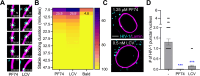
HIV-1 infection requires passage of the viral core through the nuclear pore of the cell, a process that depends on functions of the viral capsid. Recent studies have shown that HIV-1 cores enter the nucleus prior to capsid disassembly. Interactions of the viral capsid with the nuclear pore complex are necessary but not sufficient for nuclear entry, and the mechanism by which the viral core traverses the comparably sized nuclear pore is unknown. Here we show that the HIV-1 core is highly elastic and that this property is linked to nuclear entry and infectivity. Using atomic force microscopy-based approaches, we found that purified wild type cores rapidly returned to their normal conical morphology following a severe compression. Results from independently performed molecular dynamic simulations of the mature HIV-1 capsid also revealed its elastic property. Analysis of four HIV-1 capsid mutants that exhibit impaired nuclear entry revealed that the mutant viral cores are brittle. Adaptation of two of the mutant viruses in cell culture resulted in additional substitutions that restored elasticity and rescued infectivity and nuclear entry. We also show that capsid-targeting compound PF74 and the antiviral drug Lenacapavir reduce core elasticity and block HIV-1 nuclear entry at concentrations that preserve interactions between the viral core and the nuclear envelope. Our results indicate that elasticity is a fundamental property of the HIV-1 core that enables nuclear entry, thereby facilitating infection. These results provide new insights into the role of the capsid in HIV-1 nuclear entry and the antiviral mechanisms of HIV-1 capsid inhibitors.
Copyright: © 2024 Deshpande et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Update of
-
Elasticity of the HIV-1 Core Facilitates Nuclear Entry and Infection.bioRxiv [Preprint]. 2023 Sep 30:2023.09.29.560083. doi: 10.1101/2023.09.29.560083. bioRxiv. 2023. Update in: PLoS Pathog. 2024 Sep 11;20(9):e1012537. doi: 10.1371/journal.ppat.1012537. PMID: 37808653 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

