Combining short- and long-read sequencing unveils geographically structured diversity in Borrelia miyamotoi
- PMID: 39262806
- PMCID: PMC11388275
- DOI: 10.1016/j.isci.2024.110616
Combining short- and long-read sequencing unveils geographically structured diversity in Borrelia miyamotoi
Abstract
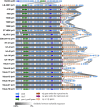
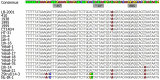
Borrelia miyamotoi is an emerging Ixodes tick-borne human pathogen in the Northern hemisphere. The aim of the current study was to compare whole genome sequences of B. miyamotoi isolates from different continents. Using a combination of Illumina and PacBio platforms and a novel genome assembly and plasmid typing pipeline, we reveal that the 21 sequenced B. miyamotoi isolates and publically available B. miyamotoi genomes from North America, Asia, and Europe form genetically distinct populations and cluster according to their geographical origin, where distinct Ixodes species are endemic. We identified 20 linear and 17 circular plasmid types and the presence of specific plasmids for isolates originating from different continents. Linear plasmids lp12, lp23, lp41, and lp72 were core plasmids found in all isolates, with lp41 consistently containing the vmp expression site. Our data provide insights into the genetic basis of vector competence, virulence, and pathogenesis of B. miyamotoi.
Keywords: Bacteriology; Geography; Microbial genomics; Phylogenetics; Techniques in genetics.
© 2024 The Authors.
Conflict of interest statement
All authors declare no competing interests.
Figures





References
Grants and funding
LinkOut - more resources
Full Text Sources

