Lysosomal TFEB-TRPML1 Axis in Astrocytes Modulates Depressive-like Behaviors
- PMID: 39264289
- PMCID: PMC11538709
- DOI: 10.1002/advs.202403389
Lysosomal TFEB-TRPML1 Axis in Astrocytes Modulates Depressive-like Behaviors
Abstract
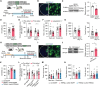
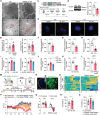
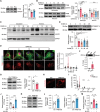
Lysosomes are important cellular structures for human health as centers for recycling, signaling, metabolism and stress adaptation. However, the potential role of lysosomes in stress-related emotions has long been overlooked. Here, it is found that lysosomal morphology in astrocytes is altered in the medial prefrontal cortex (mPFC) of susceptible mice after chronic social defeat stress. A screen of lysosome-related genes revealed that the expression of the mucolipin 1 gene (Mcoln1; protein: mucolipin TRP channel 1) is decreased in susceptible mice and depressed patients. Astrocyte-specific knockout of mucolipin TRP channel 1 (TRPML1) induced depressive-like behaviors by inhibiting lysosomal exocytosis-mediated adenosine 5'-triphosphate (ATP) release. Furthermore, this stress response of astrocytic lysosomes is mediated by the transcription factor EB (TFEB), and overexpression of TRPML1 rescued depressive-like behaviors induced by astrocyte-specific knockout of TFEB. Collectively, these findings reveal a lysosomal stress-sensing signaling pathway contributing to the development of depression and identify the lysosome as a potential target organelle for antidepressants.
Keywords: ATP; astrocytes; depressive‐like behaviors; lysosomes; mucolipin TRP channel 1; transcription factor EB.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Malhi G. S., Mann J. J., Lancet. 2018, 392, 2299. - PubMed
-
- Hammen C., Annual Rev. Clin. Psychol. 2005, 1, 293. - PubMed
-
- Leng L., Zhuang K., Liu Z., Huang C., Gao Y., Chen G., Lin H., Hu Y., Wu D., Shi M., Xie W., Sun H., Shao Z., Li H., Zhang K., Mo W., Huang T. Y., Xue M., Yuan Z., Zhang X., Bu G., Xu H., Xu Q., Zhang J., Neuron 2018, 100, 551. - PubMed
MeSH terms
Substances
Grants and funding
- 32271062/National Natural Science Foundation of China
- 2022ZD0204700/STI2030-Major Projects
- 202206060001/Guangzhou Key Research Program on Brain Science
- 2018B030334001/Key-Area Research and Development Program of Guangdong Province
- 2018B030340001/Key-Area Research and Development Program of Guangdong Province
LinkOut - more resources
Full Text Sources
Medical
