ImputeHiFI: An Imputation Method for Multiplexed DNA FISH Data by Utilizing Single-Cell Hi-C and RNA FISH Data
- PMID: 39264290
- PMCID: PMC11558076
- DOI: 10.1002/advs.202406364
ImputeHiFI: An Imputation Method for Multiplexed DNA FISH Data by Utilizing Single-Cell Hi-C and RNA FISH Data
Abstract
Although multiplexed DNA fluorescence in situ hybridization (FISH) enables tracking the spatial localization of thousands of genomic loci using probes within individual cells, the high rates of undetected probes impede the depiction of 3D chromosome structures. Current data imputation methods neither utilize single-cell Hi-C data, which elucidate 3D genome architectures using sequencing nor leverage multimodal RNA FISH data that reflect cell-type information, limiting the effectiveness of these methods in complex tissues such as the mouse brain. To this end, a novel multiplexed DNA FISH imputation method named ImputeHiFI is proposed, which fully utilizes the complementary structural information from single-cell Hi-C data and the cell type signature from RNA FISH data to obtain a high-fidelity and complete spatial location of chromatin loci. ImputeHiFI enhances cell clustering, compartment identification, and cell subtype detection at the single-cell level in the mouse brain. ImputeHiFI improves the recognition of cell-type-specific loops in three high-resolution datasets. In short, ImputeHiFI is a powerful tool capable of imputing multiplexed DNA FISH data from various resolutions and imaging protocols, facilitating studies of 3D genome structures and functions.
Keywords: 3D genomes; DNA FISH data; imputation; multimodal imaging; single‐cell Hi‐C data.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
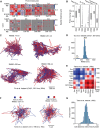





References
-
- Luo X., Liu Y., Dang D., Hu T., Hou Y., Meng X., Zhang F., Li T., Wang C., Li M., Cell 2021, 184, 723. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
