Subfunctionalization of NRC3 altered the genetic structure of the Nicotiana NRC network
- PMID: 39264953
- PMCID: PMC11421798
- DOI: 10.1371/journal.pgen.1011402
Subfunctionalization of NRC3 altered the genetic structure of the Nicotiana NRC network
Abstract
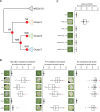
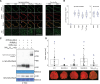
Nucleotide-binding domain and leucine-rich repeat (NLR) proteins play crucial roles in immunity against pathogens in both animals and plants. In solanaceous plants, activation of several sensor NLRs triggers their helper NLRs, known as NLR-required for cell death (NRC), to form resistosome complexes to initiate immune responses. While the sensor NLRs and downstream NRC helpers display diverse genetic compatibility, molecular evolutionary events leading to the complex network architecture remained elusive. Here, we showed that solanaceous NRC3 variants underwent subfunctionalization after the divergence of Solanum and Nicotiana, altering the genetic architecture of the NRC network in Nicotiana. Natural solanaceous NRC3 variants form three allelic groups displaying distinct compatibilities with the sensor NLR Rpi-blb2. Ancestral sequence reconstruction and analyses of natural and chimeric variants identified six key amino acids involved in sensor-helper compatibility. These residues are positioned on multiple surfaces of the resting NRC3 homodimer, collectively contributing to their compatibility with Rpi-blb2. Upon activation, Rpi-blb2-compatible NRC3 variants form membrane-associated punctate and high molecular weight complexes, and confer resistance to the late blight pathogen Phytophthora infestans. Our findings revealed how mutations in NRC alleles lead to subfunctionalization, altering sensor-helper compatibility and contributing to the increased complexity of the NRC network.
Copyright: © 2024 Huang et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
I have read the journal’s policy and the authors of this manuscript have the following competing interests: JK received funding from industry on NLR biology at the time of the study. JK has filed patents on NLR biology. Other authors have declared that no competing interests exist.
Figures




References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

