A miR-383-5p Signaling Hub Coordinates the Axon Regeneration Response to Inflammation
- PMID: 39266301
- PMCID: PMC11529811
- DOI: 10.1523/JNEUROSCI.1822-23.2024
A miR-383-5p Signaling Hub Coordinates the Axon Regeneration Response to Inflammation
Abstract
Neuroinflammation can positively influence axon regeneration following injury in the central nervous system. Inflammation promotes the release of neurotrophic molecules and stimulates intrinsic proregenerative molecular machinery in neurons, but the detailed mechanisms driving this effect are not fully understood. We evaluated how microRNAs are regulated in retinal neurons in response to intraocular inflammation to identify their potential role in axon regeneration. We found that miR-383-5p is downregulated in retinal ganglion cells in response to zymosan-induced intraocular inflammation. MiR-383-5p downregulation in neurons is sufficient to promote axon growth in vitro, and the intravitreal injection of a miR-383-5p inhibitor into the eye promotes axon regeneration following optic nerve crush. MiR-383-5p directly targets ciliary neurotrophic factor (CNTF) receptor components, and miR-383-5p inhibition sensitizes adult retinal neurons to the outgrowth-promoting effects of CNTF. Interestingly, we also demonstrate that CNTF treatment is sufficient to reduce miR-383-5p levels in neurons, constituting a positive-feedback module, whereby initial CNTF treatment reduces miR-383-5p levels, which then disinhibits CNTF receptor components to sensitize neurons to the ligand. Additionally, miR-383-5p inhibition derepresses the mitochondrial antioxidant protein peroxiredoxin-3 (PRDX3) which was required for the proregenerative effects associated with miR-383-5p loss-of-function in vitro. We have thus identified a positive-feedback mechanism that facilitates neuronal CNTF sensitivity in neurons and a new molecular signaling module that promotes inflammation-induced axon regeneration.
Keywords: axon regeneration; inflammation; microRNA; neurotrauma; neurotrophin; retina.
Copyright © 2024 the authors.
Conflict of interest statement
The authors declare no competing financial interests.
Figures





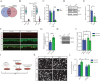

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
