POP1 Facilitates Proliferation in Triple-Negative Breast Cancer via m6A-Dependent Degradation of CDKN1A mRNA
- PMID: 39268503
- PMCID: PMC11391272
- DOI: 10.34133/research.0472
POP1 Facilitates Proliferation in Triple-Negative Breast Cancer via m6A-Dependent Degradation of CDKN1A mRNA
Abstract
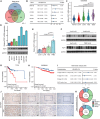
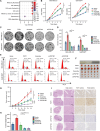
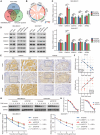
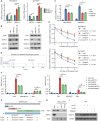
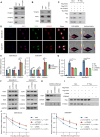
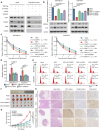
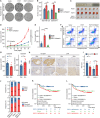
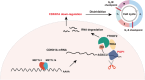
Triple-negative breast cancer (TNBC) is currently the worst prognostic subtype of breast cancer, and there is no effective treatment other than chemotherapy. Processing of precursors 1 (POP1) is the most substantially up-regulated RNA-binding protein (RBP) in TNBC. However, the role of POP1 in TNBC remains clarified. A series of molecular biological experiments in vitro and in vivo and clinical correlation analyses were conducted to clarify the biological function and regulatory mechanism of POP1 in TNBC. Here, we identified that POP1 is significantly up-regulated in TNBC and associated with poor prognosis. We further demonstrate that POP1 promotes the cell cycle and proliferation of TNBC in vitro and vivo. Mechanistically, POP1 directly binds to the coding sequence (CDS) region of CDKN1A mRNA and degrades it. The degradation process depends on the N6-methyladenosine (m6A) modification at the 497th site of CDKN1A and the recognition of this modification by YTH N6-methyladenosine RNA binding protein 2 (YTHDF2). Moreover, the m6A inhibitor STM2457 potently impaired the proliferation of POP1-overexpressed TNBC cells and improved the sensitivity to paclitaxel. In summary, our findings reveal the pivotal role of POP1 in promoting TNBC proliferation by degrading the mRNA of CDKN1A and that inhibition of m6A with STM2457 is a promising therapeutic strategy for TNBC.
Copyright © 2024 Chao Zhang et al.
Conflict of interest statement
Competing interests: The authors declare that they have no competing interests.
Figures

Similar articles
-
IGF2BP3 mediates the mRNA degradation of NF1 to promote triple-negative breast cancer progression via an m6A-dependent manner.Clin Transl Med. 2023 Sep;13(9):e1427. doi: 10.1002/ctm2.1427. Clin Transl Med. 2023. PMID: 37743642 Free PMC article.
-
POP1 promotes the progression of breast cancer through maintaining telomere integrity.Carcinogenesis. 2023 May 27;44(3):252-262. doi: 10.1093/carcin/bgad017. Carcinogenesis. 2023. PMID: 37010429
-
IGF2BP2 Drives Cell Cycle Progression in Triple-Negative Breast Cancer by Recruiting EIF4A1 to Promote the m6A-Modified CDK6 Translation Initiation Process.Adv Sci (Weinh). 2024 Jan;11(1):e2305142. doi: 10.1002/advs.202305142. Epub 2023 Nov 20. Adv Sci (Weinh). 2024. PMID: 37983610 Free PMC article.
-
A ten N6-methyladenosine-related long non-coding RNAs signature predicts prognosis of triple-negative breast cancer.J Clin Lab Anal. 2021 Jun;35(6):e23779. doi: 10.1002/jcla.23779. Epub 2021 May 2. J Clin Lab Anal. 2021. PMID: 33934391 Free PMC article.
-
WFDC21P promotes triple-negative breast cancer proliferation and migration through WFDC21P/miR-628/SMAD3 axis.Front Oncol. 2022 Oct 31;12:1032850. doi: 10.3389/fonc.2022.1032850. eCollection 2022. Front Oncol. 2022. PMID: 36387210 Free PMC article.
Cited by
-
OTUB2 promotes proliferation and metastasis of triple-negative breast cancer by deubiquitinating TRAF6.Oncol Res. 2025 Apr 18;33(5):1135-1147. doi: 10.32604/or.2025.062767. eCollection 2025. Oncol Res. 2025. PMID: 40296903 Free PMC article.
-
Exploring the therapeutic potential and in vitro validation of baicalin for the treatment of triple-negative breast cancer.Front Pharmacol. 2025 Apr 28;16:1530056. doi: 10.3389/fphar.2025.1530056. eCollection 2025. Front Pharmacol. 2025. PMID: 40356970 Free PMC article.
-
DLAT promotes triple-negative breast cancer progression via YAP1 activation.Cancer Biol Ther. 2024 Dec 31;25(1):2421578. doi: 10.1080/15384047.2024.2421578. Epub 2024 Oct 26. Cancer Biol Ther. 2024. PMID: 39460738 Free PMC article.
-
ALKBH5 suppresses gastric cancer tumorigenesis and metastasis by inhibiting the translation of uncapped WRAP53 RNA isoforms in an m6A-dependent manner.Mol Cancer. 2025 Jan 15;24(1):19. doi: 10.1186/s12943-024-02223-4. Mol Cancer. 2025. PMID: 39815301 Free PMC article.
-
eNAMPT/Ac-STAT3/DIRAS2 Axis Promotes Development and Cancer Stemness in Triple-Negative Breast Cancer by Enhancing Cytokine Crosstalk Between Tumor-Associated Macrophages and Cancer Cells.Int J Biol Sci. 2025 Feb 18;21(5):2027-2047. doi: 10.7150/ijbs.103723. eCollection 2025. Int J Biol Sci. 2025. PMID: 40083697 Free PMC article.
References
-
- Bianchini G, De Angelis C, Licata L, Gianni L. Treatment landscape of triple-negative breast cancer—Expanded options, evolving needs. Nat Rev Clin Oncol. 2022;19(2):91–113. - PubMed
-
- Peng WT, Jin X, Xu XE, Yang YS, Ma D, Shao ZM, Jiang YZ. Inhibition of ACAA1 restrains proliferation and potentiates the response to CDK4/6 inhibitors in triple-negative breast cancer. Cancer Res. 2023;83(10):1711–1724. - PubMed
-
- Liang Y, Zhang H, Song X, Yang Q. Metastatic heterogeneity of breast cancer: Molecular mechanism and potential therapeutic targets. Semin Cancer Biol. 2020;60:14–27. - PubMed
-
- Hanahan D, Weinberg RA. Hallmarks of cancer: The next generation. Cell. 2011;144(5):646–674. - PubMed
LinkOut - more resources
Full Text Sources

