Transcriptomic profiles in major depressive disorder: the role of immunometabolic and cell-cycle-related pathways in depression with different levels of inflammation
- PMID: 39271754
- PMCID: PMC11919688
- DOI: 10.1038/s41380-024-02736-w
Transcriptomic profiles in major depressive disorder: the role of immunometabolic and cell-cycle-related pathways in depression with different levels of inflammation
Abstract
Transcriptomic profiles are important indicators for molecular mechanisms and pathways involved in major depressive disorder (MDD) and its different phenotypes, such as immunometabolic depression. We performed whole-transcriptome and pathway analyses on 139 individuals from the observational, case-control, BIOmarkers in DEPression (BIODEP) study, 105 with MDD and 34 controls. We divided MDD participants based on levels of inflammation, as measured by serum high-sensitivity C-reactive protein (CRP), in n = 39 'not inflamed' (CRP < 1 mg/L), n = 31 with 'elevated CRP' (1-3 mg/L), and n = 35 with 'low-grade inflammation' (>3 mg/L). We performed whole-blood RNA sequencing using Illumina NextSeq 550 and statistical analyses with the Deseq2 package for R statistics (RUV-corrected) and subsequent pathway analyses with Ingenuity Pathway Analysis. Immunometabolic pathways were activated in individuals with CRP > 1 mg/L, although surprisingly the CRP 1-3 group showed stronger immune activation than the CRP > 3 group. The main pathways identified in the comparison between CRP < 1 group and controls were cell-cycle-related, which may be protective against immunometabolic abnormalities in this 'non-inflamed' depressed group. We further divided MDD participants based on exposure and response to antidepressants (n = 47 non-responders, n = 37 responders, and n = 22 unmedicated), and identified specific immunomodulatory and neuroprotective pathways in responders (especially vs. non-responders), which could be relevant to treatment response. In further subgroup analyses, we found that the specific transcriptional profile of responders is independent of CRP levels, and that the inhibition of cell-cycle-related pathways in MDD with CRP < 1 mg/L is present only in those who are currently depressed, and not in the responders. The present study demonstrates immunometabolic and cell-cycle-related transcriptomic pathways associated with MDD and different (CRP-based and treatment-based) MDD phenotypes, while shedding light on potential molecular mechanisms that could prevent or facilitate an individual's trajectory toward immunometabolic depression and/or treatment-non-responsive depression. The recognition and integration of these mechanisms will facilitate a precision-medicine approach in MDD.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare that they have no known conflict of interest that could have appeared to influence the work reported in this paper. Ethics approval and consent to participate: All methods were performed in accordance with the relevant guidelines and regulations. All procedures were approved by an independent research ethics committee (National Research Ethics Service East of England, Cambridge Central, UK; approval number 15/EE/0092), and the study was conducted according to the Declaration of Helsinki (see also Supplementary Methods). All participants provided informed consent in writing [5].
Figures



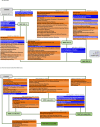
References
-
- Strawbridge R, Arnone D, Danese A, Papadopoulos A, Herane Vives A, Cleare AJ. Inflammation and clinical response to treatment in depression: a meta-analysis. Eur Neuropsychopharmacol. 2015;25:1532–43. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

