DeepASD: a deep adversarial-regularized graph learning method for ASD diagnosis with multimodal data
- PMID: 39277595
- PMCID: PMC11401938
- DOI: 10.1038/s41398-024-02972-2
DeepASD: a deep adversarial-regularized graph learning method for ASD diagnosis with multimodal data
Abstract
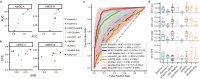
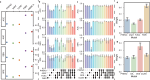
Autism Spectrum Disorder (ASD) is a prevalent neurological condition with multiple co-occurring comorbidities that seriously affect mental health. Precisely diagnosis of ASD is crucial to intervention and rehabilitation. A single modality may not fully reflect the complex mechanisms underlying ASD, and combining multiple modalities enables a more comprehensive understanding. Here, we propose, DeepASD, an end-to-end trainable regularized graph learning method for ASD prediction, which incorporates heterogeneous multimodal data and latent inter-patient relationships to better understand the pathogenesis of ASD. DeepASD first learns cross-modal feature representations through a multimodal adversarial-regularized encoder, and then constructs adaptive patient similarity networks by leveraging the representations of each modality. DeepASD exploits inter-patient relationships to boost the ASD diagnosis that is implemented by a classifier compositing of graph neural networks. We apply DeepASD to the benchmarking Autism Brain Imaging Data Exchange (ABIDE) data with four modalities. Experimental results show that the proposed DeepASD outperforms eight state-of-the-art baselines on the benchmarking ABIDE data, showing an improvement of 13.25% in accuracy, 7.69% in AUC-ROC, and 17.10% in specificity. DeepASD holds promise for a more comprehensive insight of the complex mechanisms of ASD, leading to improved diagnosis performance.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
MCBERT: A multi-modal framework for the diagnosis of autism spectrum disorder.Biol Psychol. 2025 Jan;194:108976. doi: 10.1016/j.biopsycho.2024.108976. Epub 2024 Dec 23. Biol Psychol. 2025. PMID: 39722324
-
MVS-GCN: A prior brain structure learning-guided multi-view graph convolution network for autism spectrum disorder diagnosis.Comput Biol Med. 2022 Mar;142:105239. doi: 10.1016/j.compbiomed.2022.105239. Epub 2022 Jan 19. Comput Biol Med. 2022. PMID: 35066446
-
Diagnosis of Autism Spectrum Disorders in Young Children Based on Resting-State Functional Magnetic Resonance Imaging Data Using Convolutional Neural Networks.J Digit Imaging. 2019 Dec;32(6):899-918. doi: 10.1007/s10278-019-00196-1. J Digit Imaging. 2019. PMID: 30963340 Free PMC article.
-
Do it the transformer way: A comprehensive review of brain and vision transformers for autism spectrum disorder diagnosis and classification.Comput Biol Med. 2023 Dec;167:107667. doi: 10.1016/j.compbiomed.2023.107667. Epub 2023 Nov 3. Comput Biol Med. 2023. PMID: 37939407 Review.
-
AIMAFE: Autism spectrum disorder identification with multi-atlas deep feature representation and ensemble learning.J Neurosci Methods. 2020 Sep 1;343:108840. doi: 10.1016/j.jneumeth.2020.108840. Epub 2020 Jul 9. J Neurosci Methods. 2020. PMID: 32653384 Review.
Cited by
-
Deep learning-based feature selection for detection of autism spectrum disorder.Front Artif Intell. 2025 Jun 25;8:1594372. doi: 10.3389/frai.2025.1594372. eCollection 2025. Front Artif Intell. 2025. PMID: 40636395 Free PMC article.
-
Diagnosis of Lung Cancer Using Endobronchial Ultrasonography Image Based on Multi-Scale Image and Multi-Feature Fusion Framework.Tomography. 2025 Feb 27;11(3):24. doi: 10.3390/tomography11030024. Tomography. 2025. PMID: 40137564 Free PMC article.
References
-
- Wang H, Li L, Chi L, Zhao Z. Autism screening using deep embedding representation. In: Computational science–ICCS 2019: 19th international conference, faro, Portugal, June 12–14, 2019, proceedings, part II 19. Springer; 2019. p. 160–73.
-
- Boxhoorn S, Schütz M, Mühlherr AM, Mössinger H, Luckhardt C, Freitag CM. The effect of perceptual expectation on processing gain, attention and the perceptual decision bias in children and adolescents with autism spectrum disorder (ASD). Sci Rep. 2022;12:1–14. 10.1038/s41598-022-25971-z - DOI - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

