Functional exploration and drug prediction on programmed cell death-related biomarkers in lung adenocarcinoma
- PMID: 39281570
- PMCID: PMC11401088
- DOI: 10.1016/j.heliyon.2024.e36616
Functional exploration and drug prediction on programmed cell death-related biomarkers in lung adenocarcinoma
Abstract
Background: Our study aims to perform functional exploration and drug prediction of programmed cell death (PCD)-related biomarkers in lung adenocarcinoma (LUAD).
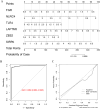
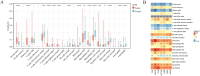
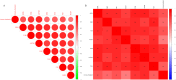
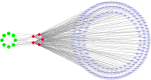
Methods: UCSC-Xena obtained LUAD-related genes. DESeq2 screened PCD-specific differentially expressed genes (DEGs), and these DEGs were intersected with genes identified by weighted gene co-expression network analysis (WGCNA) to pinpoint the key genes. KOBAS-i was used for enrichment analysis. String and GeneMania were used to construct protein interaction networks and gene-gene interaction networks, respectively. Using two machine learning algorithms to screen for key genes, and taking the intersection as biomarkers, validating via receiver operating characteristic (ROC) and in vitro experiments. Building a diagnostic model with a nomogram. Construct transcription factor (TF) regulatory network. CIBERSORT was used for immune infiltration analysis. Enrichr predicts targeted drugs and AutodockTools simulates molecular docking.
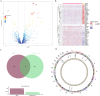
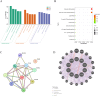
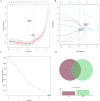
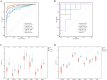
Results: 120 hub genes related to PCD were identified, and an intersection of these genes with DEGs yielded 10 key genes, which were enriched in apoptosis-related pathways. Further machine learning screening of these genes led to the selection of 7 genes, among which 6 genes (FGR, LAPTM5, SIRPA, TLR4, ZEB2, and NLRC4) exhibited significant differences upon ROC validation, ultimately serving as biomarkers, in vitro experiments also confirmed. A nomogram demonstrated their excellent diagnostic performance. These six biomarkers are correlated with the infiltration status of most immune cells, suggesting that they affect LUAD through the immune system. TF regulation analysis identified the upstream miRNAs. Finally, drug prediction yielded three potential drugs: Lenvatinib, methadone, and trimethoprim.
Conclusion: PCD-related biomarkers in LUAD were explored, which may contribute to further understanding on PCD in LUAD.
Keywords: Lung adenocarcinoma; Machine learning; Programmed cell death; Transcriptome sequencing.
© 2024 The Authors.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



References
-
- Siegel R.L., Giaquinto A.N., Jemal A. Cancer statistics, 2024. CA: a cancer journal for clinicians. 2024;74(1) - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

