Harnessing glycofluoroforms for impedimetric biosensing
- PMID: 39282644
- PMCID: PMC11393611
- DOI: 10.1039/d4sc04409f
Harnessing glycofluoroforms for impedimetric biosensing
Abstract
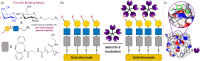
Glycans play a major role in biological cell-cell recognition and signal transduction but have found limited application in biosensors due to glycan/lectin promiscuity; multiple proteins are capable of binding to the same native glycan. Here, site-specific fluorination is used to introduce protein-glycan selectivity, and this is coupled with an electrochemical detection method to generate a novel biosensor platform. 3F-lacto-N-biose glycofluoroform is installed onto polymer tethers, which are subsequently immobilised onto gold screen printed electrodes, providing a non-fouling surface. The impedance biosensing platform is shown to selectively bind cancer-associated galectin-3 compared to control glycans and proteins. To improve the analytical capability, Bayesian statistical analysis was deployed in the equivalent circuit fitting of electrochemical impedance spectroscopy data. It is shown that Markov Chain Monte Carlo (MCMC) analysis is a helpful method for visualising experimental irreproducibility, and we apply this as a quality control step.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
The authors declare no competing financial interest.
Figures






References
-
- Thevenot D. R. Tóth K. Durst R. A. Wilson G. S. Pure Appl. Chem. 1999;71:2333–2348. doi: 10.1351/pac199971122333. - DOI
-
- Nikoleli G.-P. Siontorou C. G. Nikolelis M.-T. Bratakou S. Bendos D. K. Appl. Sci. 2019;9:1745. doi: 10.3390/app9091745. - DOI
-
- Nikoleli G.-P. Nikolelis D. P. Evtugyn G. Hianik T. TrAC, Trends Anal. Chem. 2016;79:210–221. doi: 10.1016/j.trac.2016.01.021. - DOI
-
- Blsakova A. Kveton F. Tkac J. Curr. Opin. Electrochem. 2019;14:60–65. doi: 10.1016/j.coelec.2018.12.011. - DOI
LinkOut - more resources
Full Text Sources

