Sulfated glycosaminoglycans are host epithelial cell targets of the Candida albicans toxin candidalysin
- PMID: 39285260
- PMCID: PMC11734966
- DOI: 10.1038/s41564-024-01794-8
Sulfated glycosaminoglycans are host epithelial cell targets of the Candida albicans toxin candidalysin
Abstract
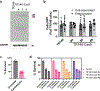
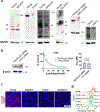
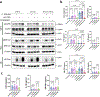
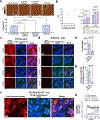
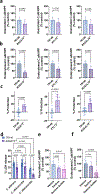
Candidalysin, a cytolytic peptide produced by the fungal pathogen Candida albicans, is a key virulence factor. However, its host cell targets remain elusive. Here we performed a genome-wide loss-of-function CRISPR screen in the TR146 human oral epithelial cell line and identified that disruption of genes (XYLT2, B3GALT6 and B3GAT3) in glycosaminoglycan (GAG) biosynthesis conferred resistance to damage induced by candidalysin and live C. albicans. Surface plasmon resonance and atomic force and electron microscopy indicated that candidalysin binds to sulfated GAGs, facilitating its enrichment on the host cell surface. Adding exogenous sulfated GAGs or the analogue dextran sulfate protected cells against candidalysin-induced damage. Dextran sulfate also inhibited C. albicans invasion and fungal-induced epithelial cell cytokine production. In mice with vulvovaginal candidiasis, topical dextran sulfate administration reduced intravaginal tissue damage and inflammation. Collectively, sulfated GAGs are epithelial cell targets of candidalysin and can be used therapeutically to protect cells from candidalysin-induced damage.
© 2024. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Competing Interests Statement
All authors declare no competing interests.
Figures



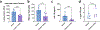




References
MeSH terms
Substances
Grants and funding
- R01 DE026600/DE/NIDCR NIH HHS/United States
- R01DE026600/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- R21AI156573/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- R01 AI134796/AI/NIAID NIH HHS/United States
- U19 AI172713/AI/NIAID NIH HHS/United States
- S10 OD028523/OD/NIH HHS/United States
- 2122027/National Science Foundation (NSF)
- U19-AI172713/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- U01-AI124319/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- R01AI134796/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- U01 AI124319/AI/NIAID NIH HHS/United States
- R35 GM140846/GM/NIGMS NIH HHS/United States
- R35GM140846/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- S10OD028523/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- R21 AI156573/AI/NIAID NIH HHS/United States

