In vivo and in vitro analyses of the role of the Prc protease in inducing mucoidy in Pseudomonas aeruginosa
- PMID: 39287400
- PMCID: PMC11500579
- DOI: 10.1128/jb.00222-24
In vivo and in vitro analyses of the role of the Prc protease in inducing mucoidy in Pseudomonas aeruginosa
Abstract
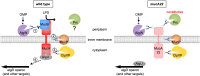
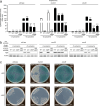
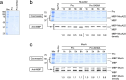
In Pseudomonas aeruginosa, alginate biosynthesis gene expression is inhibited by the transmembrane anti-sigma factor MucA, which sequesters the AlgU sigma factor. Cell envelope stress initiates cleavage of the MucA periplasmic domain by site-1 protease AlgW, followed by further MucA degradation to release AlgU. However, after colonizing the lungs of people with cystic fibrosis, P. aeruginosa converts to a mucoid form that produces alginate constitutively. Mucoid isolates often have mucA mutations, with the most common being mucA22, which truncates the periplasmic domain. MucA22 is degraded constitutively, and genetic studies suggested that the Prc protease is responsible. Some studies also suggested that Prc contributes to induction in strains with wild-type MucA, whereas others suggested the opposite. However, missing from all previous studies is a demonstration that Prc cleaves any protein directly, which leaves open the possibility that the effect of a prc null mutation is indirect. To address the ambiguities and shortfalls, we reevaluated the roles of AlgW and Prc as MucA and MucA22 site-1 proteases. In vivo analyses using three different assays and two different inducing conditions all suggested that AlgW is the only site-1 protease for wild-type MucA in any condition. In contrast, genetics suggested that AlgW or Prc act as MucA22 site-1 proteases in inducing conditions, whereas Prc is the only MucA22 site-1 protease in non-inducing conditions. For the first time, we also show that Prc is unable to degrade the periplasmic domain of wild-type MucA but does degrade the mutated periplasmic domain of MucA22 directly.
Importance: After colonizing the lungs of individuals with cystic fibrosis, Pseudomonas aeruginosa undergoes mutagenic conversion to a mucoid form, worsening the prognosis. Most mucoid isolates have a truncated negative regulatory protein MucA, which leads to constitutive production of the extracellular polysaccharide alginate. The protease Prc has been implicated, but not shown, to degrade the most common MucA variant, MucA22, to trigger alginate production. This work provides the first demonstration that the molecular mechanism of Prc involvement is direct degradation of the MucA22 periplasmic domain and perhaps other truncated MucA variants as well. MucA truncation and degradation by Prc might be the predominant mechanism of mucoid conversion in cystic fibrosis infections, suggesting that Prc activity could be a useful therapeutic target.
Keywords: Pseudomonas aeruginosa; alginate; proteases; regulation of gene expression.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Update of
-
In vivo and in vitro analysis of the role of the Prc protease in inducing mucoidy in Pseudomonas aeruginosa.bioRxiv [Preprint]. 2024 May 28:2024.05.28.596254. doi: 10.1101/2024.05.28.596254. bioRxiv. 2024. Update in: J Bacteriol. 2024 Oct 24;206(10):e0022224. doi: 10.1128/jb.00222-24. PMID: 38854061 Free PMC article. Updated. Preprint.
References
-
- Rommens JM, Iannuzzi MC, Kerem B-S, Drumm ML, Melmer G, Dean M, Rozmahel R, Cole JL, Kennedy D, Hidaka N, Zsiga M, Buchwald M, Tsui L-C, Riordan JR, Collins FS. 1989. Identification of the cystic fibrosis gene: chromosome walking and jumping. Science 245:1059–1065. doi: 10.1126/science.2772657 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

