Combined Autophagy Inhibition and Dendritic Cell Recruitment Induces Antitumor Immunity and Enhances Immune Checkpoint Blockade Sensitivity in Pancreatic Cancer
- PMID: 39288081
- PMCID: PMC11647207
- DOI: 10.1158/0008-5472.CAN-24-0830
Combined Autophagy Inhibition and Dendritic Cell Recruitment Induces Antitumor Immunity and Enhances Immune Checkpoint Blockade Sensitivity in Pancreatic Cancer
Abstract
The effect of immune checkpoint inhibitors is extremely limited in patients with pancreatic ductal adenocarcinoma (PDAC) due to the suppressive tumor immune microenvironment. Autophagy, which has been shown to play a role in antitumor immunity, has been proposed as a therapeutic target for PDAC. In this study, single-cell RNA sequencing of autophagy-deficient murine PDAC tumors revealed that autophagy inhibition in cancer cells induced dendritic cell (DC) activation. Analysis of human PDAC tumors substantiated a negative correlation between autophagy and DC activation signatures. Mechanistically, autophagy inhibition increased the intracellular accumulation of tumor antigens, which could activate DCs. Administration of chloroquine, an autophagy inhibitor, in combination with Flt3 ligand-induced DC infiltration inhibited tumor growth and increased tumor-infiltrating T lymphocytes. However, autophagy inhibition in cancer cells also induced CD8+ T-cell exhaustion with high expression of immune checkpoint LAG3. A triple-therapy comprising chloroquine, Flt3 ligand, and an anti-LAG3 antibody markedly reduced tumor growth in orthotopic syngeneic PDAC mouse models. Thus, targeting autophagy in cancer cells and activating DCs sensitize PDAC tumors to immune checkpoint inhibitor therapy, warranting further development of this treatment approach to overcome immunosuppression in pancreatic cancer. Significance: Inhibiting autophagy in pancreatic cancer cells enhances intracellular accumulation of tumor antigens to induce dendritic cell activation and synergizes with immunotherapy to markedly inhibit the growth of pancreatic ductal adenocarcinoma.
©2024 The Authors; Published by the American Association for Cancer Research.
Conflict of interest statement
No disclosures were reported.
Figures
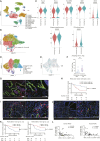
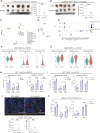

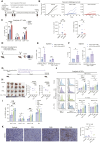
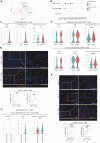
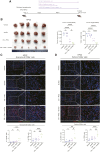
References
-
- Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA Cancer J Clin 2024;74:12–49. - PubMed
-
- Larkin J, Chiarion-Sileni V, Gonzalez R, Grob J-J, Rutkowski P, Lao CD, et al. . Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med 2019;381:1535–46. - PubMed
-
- Robert C, Schachter J, Long GV, Arance A, Grob JJ, Mortier L, et al. . Pembrolizumab versus ipilimumab in advanced melanoma. N Engl J Med 2015;372:2521–32. - PubMed
MeSH terms
Substances
Grants and funding
- JP22H00480/Japan Society for the Promotion of Science (JSPS)
- JP22H02922/Japan Society for the Promotion of Science (JSPS)
- JP22K15560/Japan Society for the Promotion of Science (JSPS)
- JP21K19530/Japan Society for the Promotion of Science (JSPS)
- JP23H02982/Japan Society for the Promotion of Science (JSPS)
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

