Solvation free energy in governing equations for DNA hybridization, protein-ligand binding, and protein folding
- PMID: 39289322
- PMCID: PMC11532980
- DOI: 10.1002/2211-5463.13897
Solvation free energy in governing equations for DNA hybridization, protein-ligand binding, and protein folding
Abstract
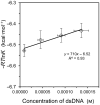
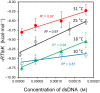
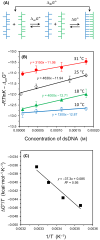
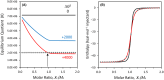
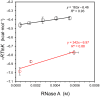
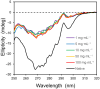
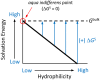
This work examines the thermodynamics of model biomolecular interactions using a governing equation that accounts for the participation of bulk water in the equilibria. In the first example, the binding affinities of two DNA duplexes, one of nine and one of 10 base pairs in length, are measured and characterized by isothermal titration calorimetry (ITC) as a function of concentration. The results indicate that the change in solvation free energy that accompanies duplex formation (ΔGS) is large and unfavorable. The duplex with the larger number of G:C pairings yields the largest change in solvation free energy, ΔGS = +460 kcal·mol-1per base pair at 25 °C. A van't Hoff analysis of the data is complicated by the varying degree of intramolecular base stacking within each DNA strand as a function of temperature. A modeling study reveals how the solvation free energy alters the output of a typical ITC experiment and leads to a good, though misleading, fit to the classical equilibrium equation. The same thermodynamic framework is applied to a model protein-ligand interaction, the binding of ribonuclease A with the nucleotide inhibitor 3'-UMP, and to a conformational equilibrium, the change in tertiary structure of α-lactalbumin in molar guanidinium chloride solutions. The ribonuclease study yields a value of ΔGS = +160 kcal·mol-1, whereas the folding equilibrium yields ΔGS ≈ 0, an apparent characteristic of hydrophobic interactions. These examples provide insight on the role of solvation energy in binding equilibria and suggest a pivot in the fundamental application of thermodynamics to solution chemistry.
Keywords: DNA duplex; bulk water; calorimetry; desolvation; hydration; ribonuclease inhibitor.
© 2024 The Author(s). FEBS Open Bio published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Thermodynamic, spectroscopic, and equilibrium binding studies of DNA sequence context effects in four 40 base pair deoxyoligonucleotides.Biochemistry. 2000 Jul 4;39(26):7835-46. doi: 10.1021/bi000326k. Biochemistry. 2000. PMID: 10869190
-
The contribution of DNA single-stranded order to the thermodynamics of duplex formation.Proc Natl Acad Sci U S A. 1991 May 1;88(9):3569-73. doi: 10.1073/pnas.88.9.3569. Proc Natl Acad Sci U S A. 1991. PMID: 2023903 Free PMC article.
-
Enthalpy and heat capacity changes for formation of an oligomeric DNA duplex: interpretation in terms of coupled processes of formation and association of single-stranded helices.Biochemistry. 1999 Jun 29;38(26):8409-22. doi: 10.1021/bi990043w. Biochemistry. 1999. PMID: 10387087
-
Stabilities of double- and triple-strand helical nucleic acids.Prog Biophys Mol Biol. 1992;58(3):225-57. doi: 10.1016/0079-6107(92)90007-s. Prog Biophys Mol Biol. 1992. PMID: 1380719 Review.
-
Differential scanning calorimetry in life science: thermodynamics, stability, molecular recognition and application in drug design.Curr Med Chem. 2005;12(17):2011-20. doi: 10.2174/0929867054546564. Curr Med Chem. 2005. PMID: 16101501 Review.
Cited by
-
Surface hydrophobic clusters modulate the folding stability and molecular recognition of the disintegrin jarastatin.J Biol Chem. 2025 Mar;301(3):108294. doi: 10.1016/j.jbc.2025.108294. Epub 2025 Feb 11. J Biol Chem. 2025. PMID: 39947470 Free PMC article.
References
-
- Eggers DK, Brewer A, Cacatian KJ, Camat LA, Castagnoli D, Chuang N, Chung LN, Do T, Huynh E, Jenpichitkulchai T et al. (2023) Model binding experiments with cucurbit[7]uril and p‐sulfonatocalix[4]arene support use of explicit solvation term in governing equation for binding equilibria. Supramol Chem 34, 94–104.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

