Activation of the NLRP1B inflammasome by caspase-8
- PMID: 39289441
- PMCID: PMC11408587
- DOI: 10.1038/s42003-024-06882-3
Activation of the NLRP1B inflammasome by caspase-8
Abstract
Cleavage of the innate immune receptor NLRP1B by various microbial proteases causes the proteasomal degradation of its N-terminal fragment and the subsequent release of a C-terminal fragment that forms an inflammasome. We reported previously that metabolic stress caused by intracellular bacteria triggers NLRP1B activation, but the mechanism by which this occurs was not elucidated. Here we demonstrate that TLR4 signaling in metabolically stressed macrophages promotes the formation of a TRIF/RIPK1/caspase-8 complex. Caspase-8 activity, induced downstream of this TLR4 pathway or through a distinct TNF receptor pathway, causes cleavage and activation of NLRP1B, which facilitates the maturation of both pro-caspase-1 and pro-caspase-8. Thus, our findings indicate that caspase-8 and NLRP1B generate a positive feedback loop that amplifies cell death processes and promotes a pro-inflammatory response through caspase-1. The ability of NLRP1B to detect caspase-8 activity suggests that this pattern recognition receptor may play a role in the defense against a variety of pathogens that induce apoptosis.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

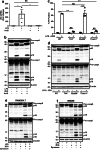






References
-
- Broz, P. & Dixit, V. M. Inflammasomes: mechanism of assembly, regulation and signalling. Nat. Rev. Immunol.16, 407–420 (2016). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

