Targeted protein relocalization via protein transport coupling
- PMID: 39294374
- PMCID: PMC11761438
- DOI: 10.1038/s41586-024-07950-8
Targeted protein relocalization via protein transport coupling
Abstract
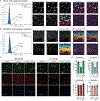
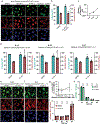
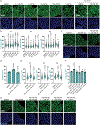
Subcellular protein localization regulates protein function and can be corrupted in cancers1 and neurodegenerative diseases2,3. The rewiring of localization to address disease-driving phenotypes would be an attractive targeted therapeutic approach. Molecules that harness the trafficking of a shuttle protein to control the subcellular localization of a target protein could enforce targeted protein relocalization and rewire the interactome. Here we identify a collection of shuttle proteins with potent ligands amenable to incorporation into targeted relocalization-activating molecules (TRAMs), and use these to relocalize endogenous proteins. Using a custom imaging analysis pipeline, we show that protein steady-state localization can be modulated through molecular coupling to shuttle proteins containing sufficiently strong localization sequences and expressed in the necessary abundance. We analyse the TRAM-induced relocalization of different proteins and then use nuclear hormone receptors as shuttles to redistribute disease-driving mutant proteins such as SMARCB1Q318X, TDP43ΔNLS and FUSR495X. TRAM-mediated relocalization of FUSR495X to the nucleus from the cytoplasm correlated with a reduction in the number of stress granules in a model of cellular stress. With methionyl aminopeptidase 2 and poly(ADP-ribose) polymerase 1 as endogenous cytoplasmic and nuclear shuttles, respectively, we demonstrate relocalization of endogenous PRMT9, SOS1 and FKBP12. Small-molecule-mediated redistribution of nicotinamide nucleotide adenylyltransferase 1 from nuclei to axons in primary neurons was able to slow axonal degeneration and pharmacologically mimic the genetic WldS gain-of-function phenotype in mice resistant to certain types of neurodegeneration4. The concept of targeted protein relocalization could therefore inspire approaches for treating disease through interactome rewiring.
© 2024. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Figures

















References
-
- Kau TR, Way JC & Silver PA Nuclear transport and cancer: from mechanism to intervention. Nat. Rev. Cancer 4, 106–117 (2004). - PubMed
-
- Hung M-C & Link W Protein localization in disease and therapy. J. Cell Sci. 124, 3381–3392 (2011). - PubMed
-
- Krauss R, Bosanac T, Devraj R, Engber T & Hughes RO Axons matter: the promise of treating neurodegenerative disorders by targeting SARM1-mediated axonal degeneration. Trends Pharmacol. Sci. 41, 281–293 (2020). - PubMed
-
- Thul PJ et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

