Utilizing machine learning algorithms for predicting risk factors for bone metastasis from right-sided colon carcinoma after complete mesocolic excision: a 10-year retrospective multicenter study
- PMID: 39298052
- PMCID: PMC11413306
- DOI: 10.1007/s12672-024-01327-z
Utilizing machine learning algorithms for predicting risk factors for bone metastasis from right-sided colon carcinoma after complete mesocolic excision: a 10-year retrospective multicenter study
Abstract
Background: Bone metastasis (BM) occurs when colon cancer cells disseminate from the primary tumor site to the skeletal system via the bloodstream or lymphatic system. The emergence of such bone metastases typically heralds a significantly poor prognosis for the patient. This study's primary aim is to develop a machine learning model to identify patients at elevated risk of bone metastasis among those with right-sided colon cancer undergoing complete mesocolonectomy (CME).
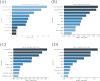
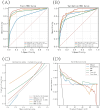
Patients and methods: The study cohort comprised 1,151 individuals diagnosed with right-sided colon cancer, with a subset of 73 patients presenting with bone metastases originating from the colon. We used univariate and multivariate regression analyses as well as four machine learning algorithms to screen variables for 38 characteristic variables such as patient demographic characteristics and surgical information. The study employed four distinct machine learning algorithms, namely, extreme gradient boosting (XGBoost), random forest (RF), support vector machine (SVM), and k-nearest neighbor algorithm (KNN), to develop the predictive model. Additionally, the model was assessed using receiver operating characteristic (ROC) curves, calibration curves, and decision curve analysis (DCA), while Shapley additive explanation (SHAP) was utilized to visualize and analyze the model.
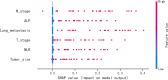
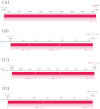
Results: The XGBoost algorithm performed the best performance among the four prediction models. In the training set, the XGBoost algorithm had an area under curve (AUC) value of 0.973 (0.953-0.994), an accuracy of 0.925 (0.913-0.936), a sensitivity of 0.921 (0.902-0.940), and a specificity of 0.908 (0.894-0.922). In the validation set, the XGBoost algorithm had an AUC value of 0.922 (0.833-0.995), an accuracy of 0.908 (0.889-0.926), a sensitivity of 0.924 (0.873-0.975), and a specificity of 0.883 (0.810-0.956). Furthermore, the AUC value of 0.83 for the external validation set suggests that the XGBoost prediction model possesses strong extrapolation capabilities. The results of SHAP analysis identified alkaline phosphatase (ALP) levels, tumor size, invasion depth, lymph node metastasis, lung metastasis, and postoperative neutrophil-to-lymphocyte ratio (NLR) levels as significant risk factors for BM from right-sided colon cancer subsequent to CME.
Conclusion: The prediction model for BM from right-sided colon cancer developed using the XGBoost machine learning algorithm in this study is both highly precise and clinically valuable.
Keywords: Bone metastasis; Colonic neoplasms; Machine learning; Prognosis; Risk factor.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Using machine learning to identify patients at high risk of developing low bone density or osteoporosis after gastrectomy: a 10-year multicenter retrospective analysis.J Cancer Res Clin Oncol. 2023 Dec;149(19):17479-17493. doi: 10.1007/s00432-023-05472-w. Epub 2023 Oct 28. J Cancer Res Clin Oncol. 2023. PMID: 37897658 Free PMC article.
-
Using machine learning algorithms to predict risk factors of heart failure after complete mesocolic excision in colorectal cancer patients.Sci Rep. 2025 Jul 15;15(1):25441. doi: 10.1038/s41598-025-11726-z. Sci Rep. 2025. PMID: 40659744 Free PMC article.
-
Identification of high-risk factors for recurrence of colon cancer following complete mesocolic excision: An 8-year retrospective study.PLoS One. 2023 Aug 11;18(8):e0289621. doi: 10.1371/journal.pone.0289621. eCollection 2023. PLoS One. 2023. PMID: 37566586 Free PMC article.
-
Ten-Year Multicenter Retrospective Study Utilizing Machine Learning Algorithms to Identify Patients at High Risk of Venous Thromboembolism After Radical Gastrectomy.Int J Gen Med. 2023 May 18;16:1909-1925. doi: 10.2147/IJGM.S408770. eCollection 2023. Int J Gen Med. 2023. PMID: 37228741 Free PMC article.
-
Interpretable machine learning model to predict surgical difficulty in laparoscopic resection for rectal cancer.Front Oncol. 2024 Feb 6;14:1337219. doi: 10.3389/fonc.2024.1337219. eCollection 2024. Front Oncol. 2024. PMID: 38380369 Free PMC article. Review.
Cited by
-
Artificial Intelligence in Orthopedic Surgery: Current Applications, Challenges, and Future Directions.MedComm (2020). 2025 Jun 25;6(7):e70260. doi: 10.1002/mco2.70260. eCollection 2025 Jul. MedComm (2020). 2025. PMID: 40567249 Free PMC article. Review.
References
-
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. 10.3322/caac.21492. - PubMed
-
- Kanthan R, Loewy J, Kanthan SC. Skeletal metastases in colorectal carcinomas: a Saskatchewan profile. Dis Colon Rectum. 1999;42:1592–7. 10.1007/bf02236213. - PubMed
-
- Rades D, Bartscht T, Janssen S, Bajrovic A, Segedin B, Schild SE. Forecasting survival probabilities after radiotherapy of metastatic epidural spinal cord compression from colorectal cancer in the elderly. Anticancer Res. 2016;36:1829–33. - PubMed
-
- Santini D, Tampellini M, Vincenzi B, Ibrahim T, Ortega C, Virzi V, Silvestris N, Berardi R, Masini C, Calipari N, et al. Natural history of bone metastasis in colorectal cancer: final results of a large Italian bone metastases study. Ann Oncol. 2012;23:2072–7. 10.1093/annonc/mdr572. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
