CYP1B1-RMDN2 Alzheimer's disease endophenotype locus identified for cerebral tau PET
- PMID: 39304655
- PMCID: PMC11415491
- DOI: 10.1038/s41467-024-52298-2
CYP1B1-RMDN2 Alzheimer's disease endophenotype locus identified for cerebral tau PET
Abstract
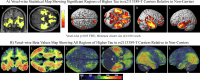
Determining the genetic architecture of Alzheimer's disease pathologies can enhance mechanistic understanding and inform precision medicine strategies. Here, we perform a genome-wide association study of cortical tau quantified by positron emission tomography in 3046 participants from 12 independent studies. The CYP1B1-RMDN2 locus is associated with tau deposition. The most significant signal is at rs2113389, explaining 4.3% of the variation in cortical tau, while APOE4 rs429358 accounts for 3.6%. rs2113389 is associated with higher tau and faster cognitive decline. Additive effects, but no interactions, are observed between rs2113389 and diagnosis, APOE4, and amyloid beta positivity. CYP1B1 expression is upregulated in AD. rs2113389 is associated with higher CYP1B1 expression and methylation levels. Mouse model studies provide additional functional evidence for a relationship between CYP1B1 and tau deposition but not amyloid beta. These results provide insight into the genetic basis of cerebral tau deposition and support novel pathways for therapeutic development in AD.
© 2024. The Author(s).
Conflict of interest statement
Dr. Apostolova received grant or other financial support from the National Institutes of Health (NIH), Alzheimer’s Association, AVID Pharmaceuticals, Life Molecular Imaging, Roche Diagnostics, and Eli Lilly. In addition, she has received consulting fees from Biogen, Two Labs, IQIVA, Florida Department of Health, Genentech, NIH Biobank, Eli Lilly, GE Healthcare, Eisai, and Roche Diagnostics. She has also received payment or honoraria from American Academy of Neurology, MillerMed, National Alzheimer’s Coordinating Center CME, CME Institute, APhA, Purdue University, Mayo Clinic, MJH Physician Education Resource, and Ohio State University. She received support for travel from the Alzheimer’s Association. She has served on Data Safety and Monitoring or Advisory Boards for IQVIA, UAB Nathan Schock Center, New Mexico Exploratory ADRC, and NIA R01 AG061111. She has a leadership role in multiple committees, including the Medical Science Council of the Alzheimer’s Association Greater Indiana Chapter, the Alzheimer’s Association Science Program Committee, and the FDA PCNS Advisory Committee. Finally, Dr. Apostolova holds stock in Cassava Neurosciences and Golden Seeds. Dr. Foroud receives support from multiple NIH grants (U24 NS095871, U24 AG021886, U24 AG056270, U01 AA026103, U10 AA008401, P30 AG010133, R01 AG019771, U01 AG032984, P30 AR072581, U01 AG057195, UL1 TR002529, U19 AG063911; U19 AG063744, U19 AG068054, R01 AG069453, U54 CA196519, R01 AG061146, R01 AG073267, R01 AG074971, U19 AG071754, R01 AG055444, R01 AG070349, U19 AG024904, R01 AG076634, U19 AG079774, U54 CA280897, U19 NS120384); the Michael J. Fox Foundation (MJFF001948); Cohen Veterans Biosciences; The Parkinson’s Disease Foundation; Children’s Tumor Foundation; Broad Institute; Lumind Foundation; and Gates Venture (0432-06-120975). Dr. Jagust has served as a consultant for Biogen, Eisai, Lilly, and Bioclinica. He has an equity interest in Optoceuticals. Aparna Vasanthakumar and Jeffrey F. Waring are employees of AbbVie and may own AbbVie stock. Dr. Hohman receives support from multiple NIH grants (U24-AG074855, P20-AG068082, R01-AG061518, R01-AG059716, R01-AG074012, RF1-AG059869). He also sits on the advisory board for Vivid Genomics and is a Senior Associate Editor for Alzheimer’s and Dementia: Translational Research and Clinical Intervention. Hyun-Sik Yang received personal fees (honorarium) from Genentech, Inc outside the submitted work. Dr. Vemuri receives funding support from the NIH. Dr. Cruchaga has received research support from: GSK and EISAI. The funders of the study had no role in the collection, analysis, or interpretation of data; in the writing of the report; or in the decision to submit the paper for publication. Dr. Cruchaga is a member of the advisory board of Vivid Genomics and Circular Genomics. Dr. Saykin receives support from multiple NIH grants (P30 AG010133, P30 AG072976, R01 AG019771, R01 AG057739, U19 AG024904, R01 LM013463, R01 AG068193, T32 AG071444, U01 AG068057, U01 AG072177 and U19 AG074879). He has also received support from Avid Radiopharmaceuticals, a subsidiary of Eli Lilly (in kind contribution of PET tracer precursor); Bayer Oncology (Scientific Advisory Board); Eisai (Scientific Advisory Board); Siemens Medical Solutions USA, Inc. (Dementia Advisory Board); NIH NHLBI (MESA Observational Study Monitoring Board); Springer-Nature Publishing (Editorial Office Support as Editor-in-Chief, Brain Imaging and Behavior). Dr. Ertekin-Taner receives support from multiple NIH grants (R01 AG061796, U19 AG074879, U01 AG046139, KL2 TR002379), Alzheimer’s Association Zenith Fellows Award and Florida Department of Health. She serves as an advisor to the Framingham Heart Study (NIH NHLBI 75N92019D00031/75N92019F00125). She also has a provisional patent application unrelated to this work. The remaining authors declare no competing interests.
Figures




References
-
- Association, A. s. 2019. Alzheimer’s disease facts and figures. Alzheimer’s & Dementia15, 321–387 (2019).
-
- Jansen, P. R. et al. Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat Genet51, 394–403 (2019). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P30 AG066444/AG/NIA NIH HHS/United States
- P50 AG016574/AG/NIA NIH HHS/United States
- U01 AG068057/AG/NIA NIH HHS/United States
- UL1 TR001108/TR/NCATS NIH HHS/United States
- R01 LM011360/LM/NLM NIH HHS/United States
- R01 AG034570/AG/NIA NIH HHS/United States
- R01 AG064877/AG/NIA NIH HHS/United States
- R01 AG041851/AG/NIA NIH HHS/United States
- U01 AG046152/AG/NIA NIH HHS/United States
- P50 GM115318/GM/NIGMS NIH HHS/United States
- R01 AG019771/AG/NIA NIH HHS/United States
- R01 AG063689/AG/NIA NIH HHS/United States
- R38 AG065762/AG/NIA NIH HHS/United States
- C06 RR018898/RR/NCRR NIH HHS/United States
- R01 AG062542/AG/NIA NIH HHS/United States
- R01 AG052521/AG/NIA NIH HHS/United States
- R01 AG056366/AG/NIA NIH HHS/United States
- W81XWH-14-2-0151/Department of Defense
- R01 AG023571/AG/NIA NIH HHS/United States
- R01 AG044546/AG/NIA NIH HHS/United States
- U01 AG072177/AG/NIA NIH HHS/United States
- P30 AG010133/AG/NIA NIH HHS/United States
- U24 AG021886/AG/NIA NIH HHS/United States
- R01-AG061796/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- R03 AG054936/AG/NIA NIH HHS/United States
- U01 AG058922/AG/NIA NIH HHS/United States
- R01 AG034676/AG/NIA NIH HHS/United States
- U01 AG061356/AG/NIA NIH HHS/United States
- U01 AG032984/AG/NIA NIH HHS/United States
- RF1 AG053303/AG/NIA NIH HHS/United States
- R37 AG057739/AG/NIA NIH HHS/United States
- U19 AG074879/AG/NIA NIH HHS/United States
- R01 AG030146/AG/NIA NIH HHS/United States
- P01 AG036694/AG/NIA NIH HHS/United States
- U01 AG024904/AG/NIA NIH HHS/United States
- U19 AG010483/AG/NIA NIH HHS/United States
- P01 AG003991/AG/NIA NIH HHS/United States
- R01 AG061518/AG/NIA NIH HHS/United States
- R01 AG039495/AG/NIA NIH HHS/United States
- P01 AG026276/AG/NIA NIH HHS/United States
- R01 AG017917/AG/NIA NIH HHS/United States
- RF1 AG058501/AG/NIA NIH HHS/United States
- U19 AG024904/AG/NIA NIH HHS/United States
- R01 AG011378/AG/NIA NIH HHS/United States
- R01 AG036042/AG/NIA NIH HHS/United States
- P30 AG010161/AG/NIA NIH HHS/United States
- R01 AG061796/AG/NIA NIH HHS/United States
- R01 AG032990/AG/NIA NIH HHS/United States
- U01 AG010483/AG/NIA NIH HHS/United States
- R01 NS080820/NS/NINDS NIH HHS/United States
- R01 AG059716/AG/NIA NIH HHS/United States
- WT101650MA/WT_/Wellcome Trust/United Kingdom
- RC2 AG036547/AG/NIA NIH HHS/United States
- R01 AG061788/AG/NIA NIH HHS/United States
- U19-AG074879/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- K23 AG062750/AG/NIA NIH HHS/United States
- U01 AG046139/AG/NIA NIH HHS/United States
- RF1 AG057473/AG/NIA NIH HHS/United States
- W81XWH-12-2-0012/Department of Defense
- R01 NS097495/NS/NINDS NIH HHS/United States
- P30 AG062677/AG/NIA NIH HHS/United States
- R01 AG052446/AG/NIA NIH HHS/United States
- K01 AG049050/AG/NIA NIH HHS/United States
- R01 AG057739/AG/NIA NIH HHS/United States
- Alzheimer's Association Zenith Fellows Award/ALZ/Alzheimer's Association/United States
- R01 AG068193/AG/NIA NIH HHS/United States
- R01 LM012535/LM/NLM NIH HHS/United States
- P01 AG003949/AG/NIA NIH HHS/United States
- U24 NS072026/NS/NINDS NIH HHS/United States
- R37 AG011378/AG/NIA NIH HHS/United States
- P30 AG066468/AG/NIA NIH HHS/United States
- P30 AG072976/AG/NIA NIH HHS/United States
- U01 AG046161/AG/NIA NIH HHS/United States
- WT_/Wellcome Trust/United Kingdom
- P30 AG019610/AG/NIA NIH HHS/United States
- R01 AG048015/AG/NIA NIH HHS/United States
- R01-NS080820/U.S. Department of Health & Human Services | NIH | National Institute of Neurological Disorders and Stroke (NINDS)
- P50 AG025711/AG/NIA NIH HHS/United States
- P01 AG017216/AG/NIA NIH HHS/United States
- P01 AG025204/AG/NIA NIH HHS/United States
- RF1 AG052525/AG/NIA NIH HHS/United States
- R01 AG018023/AG/NIA NIH HHS/United States
- U01 AG006786/AG/NIA NIH HHS/United States
- R01 AG036836/AG/NIA NIH HHS/United States
- R01 LM013463/LM/NLM NIH HHS/United States
- R01 AG015819/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

