Transcriptomic Analyses Predict Enhanced Metabolic Activity and Therapeutic Potential of mTOR Inhibitors in Acne-Prone Skin
- PMID: 39310809
- PMCID: PMC11415809
- DOI: 10.1016/j.xjidi.2024.100306
Transcriptomic Analyses Predict Enhanced Metabolic Activity and Therapeutic Potential of mTOR Inhibitors in Acne-Prone Skin
Abstract
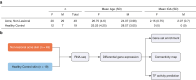
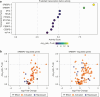
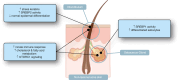
Current acne therapies center on preventing new lesions in patients with acne. These therapies were historically found to be beneficial yet were chosen without knowledge of the specific changes in the skin that favor lesion development. A major challenge in developing new treatments is the incomplete understanding of nonlesional (NL), acne-prone skin's molecular characteristics. To address this, we compared RNA-sequencing data from NL skin of 49 patients with acne (denoted as NL acne [NLA]) with those from 19 healthy controls with no acne history. We found 77 differentially expressed genes in NLA (log fold change > 1; P < .05), including genes associated with innate immunity and epidermal barrier function. Notably, K RT 6C, K RT 16, S100A8, S100A9, and lactotransferrin were upregulated, and LCE4A, LCE6A, and CTSE were downregulated. Gene set enrichment analysis revealed that metabolic pathways were enriched in NLA skin, whereas keratinization was negatively enriched. To identify compounds that could shift the gene expression signature of NLA skin toward healthy control skin, we performed connectivity mapping with the Library of Integrated Network-Based Signatures. We identified 187 compounds, particularly mTOR inhibitors, that could potentially normalize the gene expression profile of acne-prone skin to that of healthy skin. Our findings indicate that NLA skin has distinct differences in epidermal differentiation, cellular metabolism, and innate immunity that may promote lesion formation and suggest that mTOR inhibitors could restore NLA skin toward a healthier state, potentially reversing the predisposition to lesion development.
Keywords: Acne; Metabolism; Nonlesional; Transcriptomics; mTOR.
© 2024 The Authors.
Figures




Similar articles
-
Viaminate Inhibits Propionibacterium Acnes-induced Abnormal Proliferation and Keratinization of HaCat Cells by Regulating the S100A8/S100A9- MAPK Cascade.Curr Drug Targets. 2023;24(13):1055-1065. doi: 10.2174/0113894501243867230928115205. Curr Drug Targets. 2023. PMID: 37861037
-
Identification of Genes and Pathways Associated with Acne Using Integrated Bioinformatics Methods.Dermatology. 2019;235(6):445-455. doi: 10.1159/000502203. Epub 2019 Aug 27. Dermatology. 2019. PMID: 31454825
-
Steroid sulfatase activity in epidermis of acne-prone and non-acne-prone skin of patients with acne vulgaris.Arch Dermatol. 1990 Oct;126(10):1312-4. Arch Dermatol. 1990. PMID: 2145810
-
A review of skin immune processes in acne.Front Immunol. 2023 Dec 15;14:1324930. doi: 10.3389/fimmu.2023.1324930. eCollection 2023. Front Immunol. 2023. PMID: 38193084 Free PMC article. Review.
-
The Role of Skin Immune System in Acne.J Clin Med. 2022 Mar 13;11(6):1579. doi: 10.3390/jcm11061579. J Clin Med. 2022. PMID: 35329904 Free PMC article. Review.
References
-
- Aitken P., Stanescu I., Boddington L., Mahon C., Fogarasi A., Liao Y.H., et al. A novel rapamycin cream formulation improves facial angiofibromas associated with tuberous sclerosis complex: a double-blind randomized placebo-controlled trial. Br J Dermatol. 2023;189:520–530. - PubMed
-
- Arora M.K., Yadav A., Saini V. Role of hormones in acne vulgaris. Clin Biochem. 2011;44:1035–1040. - PubMed
-
- Blighe K, Lun A. PCAtools: everything principal components analysis. R package version 2.16.0. 2024. https://github.com/kevinblighe/PCAtools
LinkOut - more resources
Full Text Sources
Miscellaneous

