This is a preprint.
Plasma proteomic signatures for type 2 diabetes mellitus and related traits in the UK Biobank cohort
- PMID: 39314935
- PMCID: PMC11419213
- DOI: 10.1101/2024.09.13.24313501
Plasma proteomic signatures for type 2 diabetes mellitus and related traits in the UK Biobank cohort
Update in
-
Plasma proteomic signatures for type 2 diabetes and related traits in the UK Biobank cohort.Diabetes Res Clin Pract. 2025 Jun;224:112194. doi: 10.1016/j.diabres.2025.112194. Epub 2025 Apr 22. Diabetes Res Clin Pract. 2025. PMID: 40274105
Abstract
Aims/hypothesis: The plasma proteome holds promise as a diagnostic and prognostic tool that can accurately reflect complex human traits and disease processes. We assessed the ability of plasma proteins to predict type 2 diabetes mellitus (T2DM) and related traits.
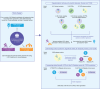
Methods: Clinical, genetic, and high-throughput proteomic data from three subcohorts of UK Biobank participants were analyzed for association with dual-energy x-ray absorptiometry (DXA) derived truncal fat (in the adiposity subcohort), estimated maximum oxygen consumption (VO2max) (in the fitness subcohort), and incident T2DM (in the T2DM subcohort). We used least absolute shrinkage and selection operator (LASSO) regression to assess the relative ability of non-proteomic and proteomic variables to associate with each trait by comparing variance explained (R2) and area under the curve (AUC) statistics between data types. Stability selection with randomized LASSO regression identified the most robustly associated proteins for each trait. The benefit of proteomic signatures (PSs) over QDiabetes, a T2DM clinical risk score, was evaluated through the derivation of delta (Δ) AUC values. We also assessed the incremental gain in model performance metrics using proteomic datasets with varying numbers of proteins. A series of two-sample Mendelian randomization (MR) analyses were conducted to identify potentially causal proteins for adiposity, fitness, and T2DM.
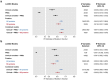
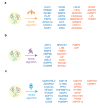
Results: Across all three subcohorts, the mean age was 56.7 years and 54.9% were female. In the T2DM subcohort, 5.8% developed incident T2DM over a median follow-up of 7.6 years. LASSO-derived PSs increased the R2 of truncal fat and VO2max over clinical and genetic factors by 0.074 and 0.057, respectively. We observed a similar improvement in T2DM prediction over the QDiabetes score [Δ AUC: 0.016 (95% CI 0.008, 0.024)] when using a robust PS derived strictly from the T2DM outcome versus a model further augmented with non-overlapping proteins associated with adiposity and fitness. A small number of proteins (29 for truncal adiposity, 18 for VO2max, and 26 for T2DM) identified by stability selection algorithms offered most of the improvement in prediction of each outcome. Filtered and clustered versions of the full proteomic dataset supplied by the UK Biobank (ranging between 600-1,500 proteins) performed comparably to the full dataset for T2DM prediction. Using MR, we identified 4 proteins as potentially causal for adiposity, 1 as potentially causal for fitness, and 4 as potentially causal for T2DM.
Conclusions/interpretation: Plasma PSs modestly improve the prediction of incident T2DM over that possible with clinical and genetic factors. Further studies are warranted to better elucidate the clinical utility of these signatures in predicting the risk of T2DM over the standard practice of using the QDiabetes score. Candidate causally associated proteins identified through MR deserve further study as potential novel therapeutic targets for T2DM.
Conflict of interest statement
Conflict of Interest None of the authors have conflicts of interest to report.
Figures


Similar articles
-
Plasma proteomic signatures for type 2 diabetes and related traits in the UK Biobank cohort.Diabetes Res Clin Pract. 2025 Jun;224:112194. doi: 10.1016/j.diabres.2025.112194. Epub 2025 Apr 22. Diabetes Res Clin Pract. 2025. PMID: 40274105
-
A plasma proteomic signature for atherosclerotic cardiovascular disease risk prediction in the UK Biobank cohort.medRxiv [Preprint]. 2024 Sep 15:2024.09.13.24313652. doi: 10.1101/2024.09.13.24313652. medRxiv. 2024. PMID: 39314942 Free PMC article. Preprint.
-
Plasma proteomic signatures of a direct measure of insulin sensitivity in two population cohorts.Diabetologia. 2023 Sep;66(9):1643-1654. doi: 10.1007/s00125-023-05946-z. Epub 2023 Jun 17. Diabetologia. 2023. PMID: 37329449 Free PMC article.
-
Development of type 2 diabetes mellitus in people with intermediate hyperglycaemia.Cochrane Database Syst Rev. 2018 Oct 29;10(10):CD012661. doi: 10.1002/14651858.CD012661.pub2. Cochrane Database Syst Rev. 2018. PMID: 30371961 Free PMC article.
-
Visceral adiposity and inflammatory bowel disease.Int J Colorectal Dis. 2021 Nov;36(11):2305-2319. doi: 10.1007/s00384-021-03968-w. Epub 2021 Jun 9. Int J Colorectal Dis. 2021. PMID: 34104989 Review.
References
-
- Einhorn D: American College of Endocrinology position statement on the insulin resistance syndrome. Endocrine practice 2003, 9:5–21. - PubMed
-
- Trout KK, Homko C, Tkacs NC: Methods of Measuring Insulin Sensitivity. Biological Research For Nursing 2007, 8(4):305–318. - PubMed
-
- Otten J, Ahrén B, Olsson T: Surrogate measures of insulin sensitivity vs the hyperinsulinaemic–euglycaemic clamp: a meta-analysis. Diabetologia 2014, 57(9):1781–1788. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
