Clinical cell-surface targets in metastatic and primary solid cancers
- PMID: 39315546
- PMCID: PMC11457844
- DOI: 10.1172/jci.insight.183674
Clinical cell-surface targets in metastatic and primary solid cancers
Abstract
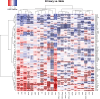
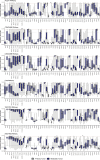
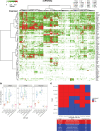
Therapies against cell-surface targets (CSTs) represent an emerging treatment class in solid malignancies. However, high-throughput investigations of CST expression across cancer types have been reliant on data sets of mostly primary tumors, despite therapeutic use most commonly in metastatic disease. We identified a total of 818 clinical trials of CST therapies with 78 CSTs. We assembled a data set spanning RNA-seq and microarrays in 7,927 benign samples, 16,866 primary tumor samples, and 6,124 metastatic tumor samples. We also utilized single-cell RNA-seq data from 36 benign tissues and 558 primary and metastatic tumor samples, and matched RNA versus protein expression in 29 benign tissue samples, 1,075 tumor samples, and 942 cell lines. High RNA expression accurately predicted high protein expression across CST therapies in benign tissues, tumor samples, and cell lines. We compared metastatic versus primary tumor expression, identified potential opportunities for repositioning, and matched cell lines to tumor types based on CST and global RNA expression. We evaluated single-cell heterogeneity across tumors, and identified rare normal cell subpopulations that may contribute to toxicity. Finally, we identified combinations of CST therapies for which bispecific approaches could improve tumor specificity. This study helps better define the landscape of CST expression in metastatic and primary cancers.
Keywords: Clinical practice; Oncology.
Figures



References
-
- National Comprehensive Cancer Network. Breast Cancer (version 1.2024). https://www.nccn.org/professionals/physician_gls/pdf/breast.pdf Accessed February 25, 2024.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

