Recycling Materials for Sustainable DNA Origami Manufacturing
- PMID: 39315689
- PMCID: PMC11451448
- DOI: 10.1021/acs.nanolett.4c02695
Recycling Materials for Sustainable DNA Origami Manufacturing
Abstract
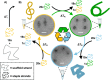
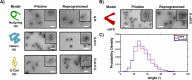
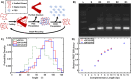
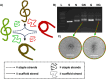
DNA origami nanotechnology has great potential in multiple fields including biomedical, biophysical, and nanofabrication applications. However, current production pipelines lead to single-use devices incorporating a small fraction of initial reactants, resulting in a wasteful manufacturing process. Here, we introduce two complementary approaches to overcome these limitations by recycling the strand components of DNA origami nanostructures (DONs). We demonstrate reprogramming entire DONs into new devices, reusing scaffold strands. We validate this approach by reprogramming DONs with complex geometries into each other, using their distinct geometries to verify successful scaffold recycling. We reprogram one DON into a dynamic structure and show both pristine and recycled structures display similar properties. Second, we demonstrate the recovery of excess staple strands postassembly and fold DONs with these recycled strands, showing these structures exhibit the expected geometry and dynamic properties. Finally, we demonstrate the combination of both approaches, successfully fabricating DONs solely from recycled DNA components.
Keywords: DNA Nanotechnology; DNA Origami; Recycling; Reprogrammable Nanomaterials; Sustainable Manufacturing; Waste Reduction.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
References
-
- Dey S.; Fan C.; Gothelf K. V.; Li J.; Lin C.; Liu L.; Liu N.; Nijenhuis M. A. D.; Saccà B.; Simmel F. C.; Yan H.; Zhan P. DNA Origami. Nat. Rev. Methods Primer 2021, 1 (1), 1–24. 10.1038/s43586-020-00009-8. - DOI
-
- Zeng Y. C.; Young O. J.; Wintersinger C. M.; Anastassacos F. M.; MacDonald J. I.; Isinelli G.; Dellacherie M. O.; Sobral M.; Bai H.; Graveline A. R.; Vernet A.; Sanchez M.; Mulligan K.; Choi Y.; Ferrante T. C.; Keskin D. B.; Fell G. G.; Neuberg D.; Wu C. J.; Mooney D. J.; Kwon I. C.; Ryu J. H.; Shih W. M. Fine Tuning of CpG Spatial Distribution with DNA Origami for Improved Cancer Vaccination. Nat. Nanotechnol. 2024, 19 (7), 1055–1065. 10.1038/s41565-024-01615-3. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

