3,N4-Etheno-5-methylcytosine blocks TET1-3 oxidation but is repaired by ALKBH2, 3 and FTO
- PMID: 39315710
- PMCID: PMC11551763
- DOI: 10.1093/nar/gkae818
3,N4-Etheno-5-methylcytosine blocks TET1-3 oxidation but is repaired by ALKBH2, 3 and FTO
Abstract
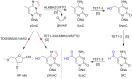
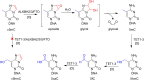
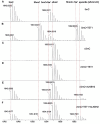
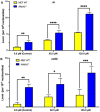
5-Methyldeoxycytidine (5mC) is a major epigenetic marker that regulates cellular functions in mammals. Endogenous lipid peroxidation can convert 5mC into 3,N4-etheno-5-methylcytosine (ϵ5mC). ϵ5mC is structurally similar to the mutagenic analog 3,N4-ethenocytosine (ϵC), which is repaired by AlkB family enzymes in the direct reversal repair (DRR) pathway and excised by DNA glycosylases in the base excision repair (BER) pathway. However, the repair of ϵ5mC has not been reported. Here, we examined the activities against ϵ5mC by DRR and BER enzymes and TET1-3, enzymes that modify the 5-methyl group in 5mC. We found that the etheno modification of 5mC blocks oxidation by TET1-3. Conversely, three human homologs in the AlkB family, ALKBH2, 3 and FTO were able to repair ϵ5mC to 5mC, which was subsequently modified by TET1 to 5-hydroxymethylcytosine. We also demonstrated that ALKBH2 likely repairs ϵ5mC in MEF cells. Another homolog, ALKBH5, could not repair ϵ5mC. Also, ϵ5mC is not a substrate for BER glycosylases SMUG1, AAG, or TDG. These findings indicate DRR committed by ALKBH2, 3 and FTO could reduce the detrimental effects of ϵ5mC in genetics and epigenetics and may work together with TET enzymes to modulate epigenetic regulations.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

Similar articles
-
DNA repair enzymes ALKBH2, ALKBH3, and AlkB oxidize 5-methylcytosine to 5-hydroxymethylcytosine, 5-formylcytosine and 5-carboxylcytosine in vitro.Nucleic Acids Res. 2019 Jun 20;47(11):5522-5529. doi: 10.1093/nar/gkz395. Nucleic Acids Res. 2019. PMID: 31114894 Free PMC article.
-
TET1 overexpression affects cell proliferation and apoptosis in aging ovaries.J Assist Reprod Genet. 2024 Dec;41(12):3491-3502. doi: 10.1007/s10815-024-03271-x. Epub 2024 Sep 25. J Assist Reprod Genet. 2024. PMID: 39317913
-
Insights into the Direct Oxidative Repair of Etheno Lesions: MD and QM/MM Study on the Substrate Scope of ALKBH2 and AlkB.DNA Repair (Amst). 2020 Dec;96:102944. doi: 10.1016/j.dnarep.2020.102944. Epub 2020 Sep 9. DNA Repair (Amst). 2020. PMID: 33161373 Free PMC article.
-
TET Enzymes in the Immune System: From DNA Demethylation to Immunotherapy, Inflammation, and Cancer.Annu Rev Immunol. 2024 Jun;42(1):455-488. doi: 10.1146/annurev-immunol-080223-044610. Epub 2024 Jun 14. Annu Rev Immunol. 2024. PMID: 38360546 Review.
-
The TET-TDG axis in T cells and biological processes.Int Immunol. 2025 May 21;37(6):299-312. doi: 10.1093/intimm/dxaf006. Int Immunol. 2025. PMID: 39921704 Review.
References
-
- Wu X., Zhang Y.. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat. Rev. Genet. 2017; 18:517–534. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

