Idiosyncratic genome evolution of the thermophilic cyanobacterium Synechococcus at the limits of phototrophy
- PMID: 39319368
- PMCID: PMC11456837
- DOI: 10.1093/ismejo/wrae184
Idiosyncratic genome evolution of the thermophilic cyanobacterium Synechococcus at the limits of phototrophy
Abstract
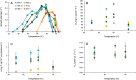
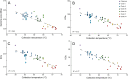
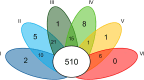
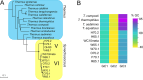
Thermophilic microorganisms are expected to have smaller cells and genomes compared with mesophiles, a higher proportion of horizontally acquired genes, and distinct nucleotide and amino acid composition signatures. Here, we took an integrative approach to investigate these apparent correlates of thermophily for Synechococcus A/B cyanobacteria, which include the most heat-tolerant phototrophs on the planet. Phylogenomics confirmed a unique origin of different thermotolerance ecotypes, with low levels of continued gene flow between ecologically divergent but overlapping populations, which has shaped the distribution of phenotypic traits along these geothermal gradients. More thermotolerant strains do have smaller genomes, but genome reduction is associated with a decrease in community richness and metabolic diversity, rather than with cell size. Horizontal gene transfer played only a limited role during Synechococcus evolution, but, the most thermotolerant strains have acquired a Thermus tRNA modification enzyme that may stabilize translation at high temperatures. Although nucleotide base composition was not associated with thermotolerance, we found a general replacement of aspartate with glutamate, as well as a dramatic remodeling of amino acid composition at the highest temperatures that substantially differed from previous predictions. We conclude that Synechococcus A/B genome diversification largely does not conform to the standard view of temperature adaptation. In addition, carbon fixation was more thermolabile than photosynthetic oxygen evolution for the most thermotolerant strains compared with less tolerant lineages. This suggests that increased flow of reducing power generated during the light reactions to an electron sink(s) beyond carbon dioxide has emerged during temperature adaptation of these bacteria.
Keywords: adaptation; cyanobacteria; genomes; horizontal gene transfer; hot springs; thermophile.
© The Author(s) 2024. Published by Oxford University Press on behalf of the International Society for Microbial Ecology.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Distribution and Genomic Variation of Thermophilic Cyanobacteria in Diverse Microbial Mats at the Upper Temperature Limits of Photosynthesis.mSystems. 2022 Oct 26;7(5):e0031722. doi: 10.1128/msystems.00317-22. Epub 2022 Aug 18. mSystems. 2022. PMID: 35980085 Free PMC article.
-
Temperature and Geographic Location Impact the Distribution and Diversity of Photoautotrophic Gene Variants in Alkaline Yellowstone Hot Springs.Microbiol Spectr. 2022 Jun 29;10(3):e0146521. doi: 10.1128/spectrum.01465-21. Epub 2022 May 16. Microbiol Spectr. 2022. PMID: 35575591 Free PMC article.
-
Photosynthetic temperature adaptation during niche diversification of the thermophilic cyanobacterium Synechococcus A/B clade.ISME J. 2017 Apr;11(4):1053-1057. doi: 10.1038/ismej.2016.173. Epub 2016 Dec 16. ISME J. 2017. PMID: 27983722 Free PMC article.
-
Ecological Divergence with Gene Flow in a Thermophilic Cyanobacterium.Microb Ecol. 2019 Jul;78(1):33-41. doi: 10.1007/s00248-018-1267-0. Epub 2018 Sep 28. Microb Ecol. 2019. PMID: 30267129
-
[High-temperature adaptation mechanisms and biotechnological potentials of thermophilic cyanobacteria].Sheng Wu Gong Cheng Xue Bao. 2024 Jun 25;40(6):1752-1775. doi: 10.13345/j.cjb.230645. Sheng Wu Gong Cheng Xue Bao. 2024. PMID: 38914490 Review. Chinese.
Cited by
-
Comparative genomics of thermosynechococcaceae and thermostichaceae: insights into codon usage bias.Acta Biochim Pol. 2025 Jan 8;71:13825. doi: 10.3389/abp.2024.13825. eCollection 2024. Acta Biochim Pol. 2025. PMID: 39845100 Free PMC article.
-
Boundaries of photosynthesis: adaptations of carbon fixation in extreme environments.FEBS Open Bio. 2025 Jul;15(7):1028-1040. doi: 10.1002/2211-5463.70047. Epub 2025 May 19. FEBS Open Bio. 2025. PMID: 40388604 Free PMC article. Review.
References
-
- Hochachka PW, Somero GN. Biochemical Adaptation: Mechanism and Process in Physiological Evolution. Oxford: Oxford University Press, 2002. 10.1093/oso/9780195117028.001.0001. - DOI
-
- Angilletta MJ. Thermal Adaptation: A Theoretical and Empirical Synthesis. Oxford: Oxford University Press, 2009, 10.1093/acprof:oso/9780198570875.001.1. - DOI
-
- Shuter BJ, Thomas JE, Taylor WDet al. . Phenotypic correlates of genomic DNA content in unicellular eukaryotes and other cells. Am Nat 1983;122:26–44. 10.1086/284116 - DOI
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

