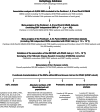
Polymorphisms within autophagy-related genes as susceptibility biomarkers for pancreatic cancer: A meta-analysis of three large European cohorts and functional characterization
- PMID: 39319538
- PMCID: PMC11578083
- DOI: 10.1002/ijc.35196
Polymorphisms within autophagy-related genes as susceptibility biomarkers for pancreatic cancer: A meta-analysis of three large European cohorts and functional characterization
Erratum in
-
Correction to "Polymorphisms within autophagy-related genes as susceptibility biomarkers for pancreatic cancer: A meta-analysis of three large European cohorts and functional characterization".Int J Cancer. 2025 Sep 1;157(5):1022. doi: 10.1002/ijc.35510. Epub 2025 Jun 6. Int J Cancer. 2025. PMID: 40480799 Free PMC article. No abstract available.
Abstract
Pancreatic ductal adenocarcinoma (PDAC) is one of the most lethal cancers with patients having unresectable or metastatic disease at diagnosis, with poor prognosis and very short survival. Given that genetic variation within autophagy-related genes influences autophagic flux and susceptibility to solid cancers, we decided to investigate whether 55,583 single nucleotide polymorphisms (SNPs) within 234 autophagy-related genes could influence the risk of developing PDAC in three large independent cohorts of European ancestry including 12,754 PDAC cases and 324,926 controls. The meta-analysis of these populations identified, for the first time, the association of the BIDrs9604789 variant with an increased risk of developing the disease (ORMeta = 1.31, p = 9.67 × 10-6). We also confirmed the association of TP63rs1515496 and TP63rs35389543 variants with PDAC risk (OR = 0.89, p = 6.27 × 10-8 and OR = 1.16, p = 2.74 × 10-5). Although it is known that BID induces autophagy and TP63 promotes cell growth, cell motility and invasion, we also found that carriers of the TP63rs1515496G allele had increased numbers of FOXP3+ Helios+ T regulatory cells and CD45RA+ T regulatory cells (p = 7.67 × 10-4 and p = 1.56 × 10-3), but also decreased levels of CD4+ T regulatory cells (p = 7.86 × 10-4). These results were in agreement with research suggesting that the TP63rs1515496 variant alters binding sites for FOXA1 and CTCF, which are transcription factors involved in modulating specific subsets of regulatory T cells. In conclusion, this study identifies BID as new susceptibility locus for PDAC and confirms previous studies suggesting that the TP63 gene is involved in the development of PDAC. This study also suggests new pathogenic mechanisms of the TP63 locus in PDAC.
Keywords: autophagy; functional characterization; genetic variants; pancreatic cancer; polymorphisms; susceptibility.
© 2024 The Author(s). International Journal of Cancer published by John Wiley & Sons Ltd on behalf of UICC.
Conflict of interest statement
All authors have no competing interests to disclose.
Figures
References
-
- Kleeff J, Korc M, Apte M, et al. Pancreatic cancer. Nat Rev Dis Primers. 2016;2:16022. - PubMed
-
- Ryan DP, Hong TS, Bardeesy N. Pancreatic adenocarcinoma. N Engl J Med. 2014;371:2140‐2141. - PubMed
-
- De La Cruz MS, Young AP, Ruffin MT. Diagnosis and management of pancreatic cancer. Am Fam Physician. 2014;89:626‐632. - PubMed
-
- Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17‐48. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- GACR21-27902 and AZVNU21-03-00145/Czech Republic Ministry of Health
- "Casa Sollievo della Sofferenza" Hospital, San Giovanni Rotondo
- PY20/01282/Consejería de Transformación Económica, Industria, Conocimiento y Universidades and FEDER
- PI17/02256/Instituto de Salud Carlos III
- PI20/01845/Instituto de Salud Carlos III
LinkOut - more resources
Full Text Sources
Medical
Research Materials