The Dsc complex and its role in Golgi quality control
- PMID: 39324639
- PMCID: PMC11555709
- DOI: 10.1042/BST20230375
The Dsc complex and its role in Golgi quality control
Abstract
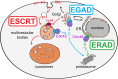
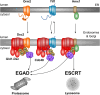
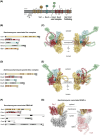
Membrane proteins play crucial roles in cellular functions. However, processes such as the insertion of membrane proteins into the endoplasmic reticulum (ER), their folding into native structures, the assembly of multi-subunit membrane protein complexes, and their targeting from the ER to specific organelles are prone to errors and have a relatively high failure rate. To prevent the accumulation of defective or orphaned membrane proteins, quality control mechanisms assess folding, quantity, and localization of these proteins. This quality control is vital for preserving organelle integrity and maintaining cellular health. In this mini-review, we will focus on how selective membrane protein quality control at the Golgi apparatus, particularly through the defective for SREBP cleavage (Dsc) ubiquitin ligase complex, detects orphaned proteins and prevents their mis-localization to other organelles.
Keywords: Golgi; quality control; ubiquitin.
© 2024 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures

Similar articles
-
The Dsc ubiquitin ligase complex identifies transmembrane degrons to degrade orphaned proteins at the Golgi.Nat Commun. 2024 Oct 26;15(1):9257. doi: 10.1038/s41467-024-53676-6. Nat Commun. 2024. PMID: 39461958 Free PMC article.
-
Endoplasmic Reticulum Exit of Golgi-resident Defective for SREBP Cleavage (Dsc) E3 Ligase Complex Requires Its Activity.J Biol Chem. 2015 Jun 5;290(23):14430-40. doi: 10.1074/jbc.M114.630863. Epub 2015 Apr 27. J Biol Chem. 2015. PMID: 25918164 Free PMC article.
-
The sterol-sensing endoplasmic reticulum (ER) membrane protein TRC8 hampers ER to Golgi transport of sterol regulatory element-binding protein-2 (SREBP-2)/SREBP cleavage-activated protein and reduces SREBP-2 cleavage.J Biol Chem. 2009 Oct 16;284(42):28995-9004. doi: 10.1074/jbc.M109.041376. Epub 2009 Aug 25. J Biol Chem. 2009. PMID: 19706601 Free PMC article.
-
Protein quality control at the Golgi.Curr Opin Cell Biol. 2022 Apr;75:102074. doi: 10.1016/j.ceb.2022.02.008. Epub 2022 Mar 29. Curr Opin Cell Biol. 2022. PMID: 35364487 Review.
-
How Is the Fidelity of Proteins Ensured in Terms of Both Quality and Quantity at the Endoplasmic Reticulum? Mechanistic Insights into E3 Ubiquitin Ligases.Int J Mol Sci. 2021 Feb 19;22(4):2078. doi: 10.3390/ijms22042078. Int J Mol Sci. 2021. PMID: 33669844 Free PMC article. Review.
Cited by
-
Lysosomal quality control Review.Autophagy. 2025 Jul;21(7):1413-1432. doi: 10.1080/15548627.2025.2469206. Epub 2025 Feb 24. Autophagy. 2025. PMID: 39968899 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

