ReBiA-Robotic Enabled Biological Automation: 3D Epithelial Tissue Production
- PMID: 39324843
- PMCID: PMC11615785
- DOI: 10.1002/advs.202406608
ReBiA-Robotic Enabled Biological Automation: 3D Epithelial Tissue Production
Abstract
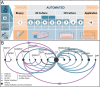
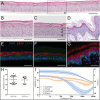
The Food and Drug Administration's recent decision to eliminate mandatory animal testing for drug approval marks a significant shift to alternative methods. Similarly, the European Parliament is advocating for a faster transition, reflecting public preference for animal-free research practices. In vitro tissue models are increasingly recognized as valuable tools for regulatory assessments before clinical trials, in line with the 3R principles (Replace, Reduce, Refine). Despite their potential, barriers such as the need for standardization, availability, and cost hinder their widespread adoption. To address these challenges, the Robotic Enabled Biological Automation (ReBiA) system is developed. This system uses a dual-arm robot capable of standardizing laboratory processes within a closed automated environment, translating manual processes into automated ones. This reduces the need for process-specific developments, making in vitro tissue models more consistent and cost-effective. ReBiA's performance is demonstrated through producing human reconstructed epidermis, human airway epithelial models, and human intestinal organoids. Analyses confirm that these models match the morphology and protein expression of manually prepared and native tissues, with similar cell viability. These successes highlight ReBiA's potential to lower barriers to broader adoption of in vitro tissue models, supporting a shift toward more ethical and advanced research methods.
Keywords: 3R; AI tools; alternatives to animal testing; lab automation; tissue engineering.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
