Far-red light photoacclimation in a desert Chroococcidiopsis strain with a reduced FaRLiP gene cluster and expression of its chlorophyll f synthase in space-resistant isolates
- PMID: 39328908
- PMCID: PMC11424453
- DOI: 10.3389/fmicb.2024.1450575
Far-red light photoacclimation in a desert Chroococcidiopsis strain with a reduced FaRLiP gene cluster and expression of its chlorophyll f synthase in space-resistant isolates
Abstract
Introduction: Some cyanobacteria can use far-red light (FRL) to drive oxygenic photosynthesis, a phenomenon known as Far-Red Light Photoacclimation (FaRLiP). It can expand photosynthetically active radiation beyond the visible light (VL) range. Therefore, it holds promise for biotechnological applications and may prove useful for the future human exploration of outer space. Typically, FaRLiP relies on a cluster of ~20 genes, encoding paralogs of the standard photosynthetic machinery. One of them, a highly divergent D1 gene known as chlF (or psbA4), is the synthase responsible for the formation of the FRL-absorbing chlorophyll f (Chl f) that is essential for FaRLiP. The minimum gene set required for this phenotype is unclear. The desert cyanobacterium Chroococcidiopsis sp. CCMEE 010 is unusual in being capable of FaRLiP with a reduced gene cluster (15 genes), and it lacks most of the genes encoding FR-Photosystem I.
Methods: Here we investigated whether the reduced gene cluster of Chroococcidiopsis sp. CCMEE 010 is transcriptionally regulated by FRL and characterized the spectral changes that occur during the FaRLiP response of Chroococcidiopsis sp. CCMEE 010. In addition, the heterologous expression of the Chl f synthase from CCMEE 010 was attempted in three closely related desert strains of Chroococcidiopsis.
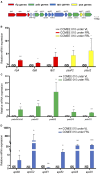
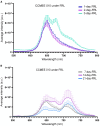
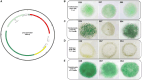
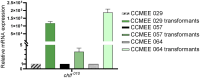
Results: All 15 genes of the FaRLiP cluster were preferentially expressed under FRL, accompanied by a progressive red-shift of the photosynthetic absorption spectrum. The Chl f synthase from CCMEE 010 was successfully expressed in two desert strains of Chroococcidiopsis and transformants could be selected in both VL and FRL.
Discussion: In Chroococcidiopsis sp. CCME 010, all the far-red genes of the unusually reduced FaRLiP cluster, are transcriptionally regulated by FRL and two closely related desert strains heterologously expressing the chlF010 gene could grow in FRL. Since the transformation hosts had been reported to survive outer space conditions, such an achievement lays the foundation toward novel cyanobacteria-based technologies to support human space exploration.
Keywords: Chl f synthase; Chroococcidiopsis; FaRLiP; genetic manipulation; space exploration.
Copyright © 2024 di Stefano, Battistuzzi, La Rocca, Selinger, Nürnberg and Billi.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures

Similar articles
-
Identification of far-red light acclimation in an endolithic Chroococcidiopsis strain and associated genomic features: Implications for oxygenic photosynthesis on exoplanets.Front Microbiol. 2022 Aug 4;13:933404. doi: 10.3389/fmicb.2022.933404. eCollection 2022. Front Microbiol. 2022. PMID: 35992689 Free PMC article.
-
Characterization of cyanobacterial allophycocyanins absorbing far-red light.Photosynth Res. 2020 Sep;145(3):189-207. doi: 10.1007/s11120-020-00775-2. Epub 2020 Jul 24. Photosynth Res. 2020. PMID: 32710194
-
RfpA, RfpB, and RfpC are the Master Control Elements of Far-Red Light Photoacclimation (FaRLiP).Front Microbiol. 2015 Nov 25;6:1303. doi: 10.3389/fmicb.2015.01303. eCollection 2015. Front Microbiol. 2015. PMID: 26635768 Free PMC article.
-
Adaptive and acclimative responses of cyanobacteria to far-red light.Environ Microbiol. 2015 Oct;17(10):3450-65. doi: 10.1111/1462-2920.12992. Epub 2015 Aug 27. Environ Microbiol. 2015. PMID: 26234306 Review.
-
Adaptation processes in Halomicronema hongdechloris, an example of the light-induced optimization of the photosynthetic apparatus on hierarchical time scales.Front Plant Sci. 2024 Jul 10;15:1359195. doi: 10.3389/fpls.2024.1359195. eCollection 2024. Front Plant Sci. 2024. PMID: 39049856 Free PMC article. Review.
References
-
- Antonaru L. A., Selinger V. M., Jung P., Di Stefano G., Sanderson N. D., Barker L., et al. . (2023). Common loss of far-red light photoacclimation in cyanobacteria from hot and cold deserts: a case study in the Chroococcidiopsidales. ISME Commun. 3:113. 10.1038/s43705-023-00319-4 - DOI - PMC - PubMed
-
- Averina S., Velichko N., Senatskaya E., Pinevich A. (2018). Far-red light photoadaptations in aquatic cyanobacteria. Hydrobiologia 813, 1–17. 10.1007/s10750-018-3519-x - DOI
-
- Battistuzzi M., Cocola L., Claudi R., Pozzer A. C., Segalla A., Simionato D., et al. . (2023). Oxygenic photosynthetic responses of cyanobacteria exposed under an M-dwarf starlight simulator: implications for exoplanet's habitability. Front. Plant Sci. 14:1070359. 10.3389/fpls.2023.1070359 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

