Chagas Disease Diagnosis with Trypanosoma cruzi-Exclusive Epitopes in GFP
- PMID: 39340059
- PMCID: PMC11435546
- DOI: 10.3390/vaccines12091029
Chagas Disease Diagnosis with Trypanosoma cruzi-Exclusive Epitopes in GFP
Abstract
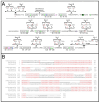
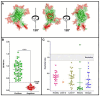
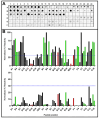
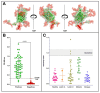
Serological tests are critical tools in the fight against infectious disease. They detect antibodies produced during an adaptive immune response against a pathogen with an immunological reagent, whose antibody binding characteristics define the specificity and sensitivity of the assay. While pathogen proteins have conveniently served as reagents, their performance is limited by the natural grouping of specific and non-specific antibody binding sites, epitopes. An attractive solution is to build synthetic proteins that only contains pathogen-specific epitopes, which could theoretically reach 100% specificity. However, the genesis of de novo proteins remains a challenge. To address the uncertainty of producing a synthetic protein, we have repurposed the beta barrel of fluorescent proteins into a receptacle that can receive several epitope sequences without compromising its ability to be expressed. Here, two versions of a multiepitope protein were built using the receptacle that differ by their grouping of epitopes specific to the parasite Trypanosoma cruzi, the causative agent for Chagas disease. An evaluation of their performance as the capture reagent in ELISAs showed near-complete agreement with recommended diagnostic protocols. The results suggest that a single assay could be developed for the diagnosis of Chagas disease and that this approach could be applied to other diseases.
Keywords: Chagas disease; ELISA; Green Fluorescent Protein; Serodiagnosis; Trypanosoma cruzi.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
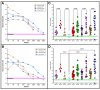
Similar articles
-
Chagas disease-specific antigens: characterization of epitopes in CRA/FRA by synthetic peptide mapping and evaluation by ELISA-peptide assay.BMC Infect Dis. 2013 Dec 3;13:568. doi: 10.1186/1471-2334-13-568. BMC Infect Dis. 2013. PMID: 24299278 Free PMC article.
-
Next-generation ELISA diagnostic assay for Chagas Disease based on the combination of short peptidic epitopes.PLoS Negl Trop Dis. 2017 Oct 9;11(10):e0005972. doi: 10.1371/journal.pntd.0005972. eCollection 2017 Oct. PLoS Negl Trop Dis. 2017. PMID: 28991925 Free PMC article.
-
Identification of highly conserved Trypanosoma cruzi antigens for the development of a universal serological diagnostic assay.Emerg Microbes Infect. 2024 Dec;13(1):2315964. doi: 10.1080/22221751.2024.2315964. Epub 2024 Feb 21. Emerg Microbes Infect. 2024. PMID: 38381980 Free PMC article.
-
Trypanosoma cruzi-specific CD8+ T cells and other immunological hallmarks in chronic Chagas cardiomyopathy: Two decades of research.Front Cell Infect Microbiol. 2023 Jan 4;12:1075717. doi: 10.3389/fcimb.2022.1075717. eCollection 2022. Front Cell Infect Microbiol. 2023. PMID: 36683674 Free PMC article. Review.
-
Trypanosoma cruzi genetic diversity: Something new for something known about Chagas disease manifestations, serodiagnosis and drug sensitivity.Acta Trop. 2018 Aug;184:38-52. doi: 10.1016/j.actatropica.2017.09.017. Epub 2017 Sep 21. Acta Trop. 2018. PMID: 28941731 Review.
References
-
- Chagas C. Nova tripanossomíase humana. Mem. Inst. Oswaldo Cruz. 1909;1:159–219. doi: 10.1590/S0074-02761909000200008. - DOI
-
- Hotez P.J., Bottazzi M.E., Franco-Paredes C., Ault S.K., Periago M.R. The neglected tropical diseases of Latin America and the Caribbean: A review of disease burden and distribution and a roadmap for control and elimination. PLoS Negl. Trop. Dis. 2008;2:e300. doi: 10.1371/journal.pntd.0000300. - DOI - PMC - PubMed
-
- WHO Chagas Disease (American Trypanosomiasis) 2024. [(accessed on 7 April 2024)]. Available online: https://www.who.int/en/news-room/fact-sheets/detail/chagas-disease-(amer...
Grants and funding
- (#SEI26003/01279/2021/Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro
- 200.960/2022/Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro
- 305157-2020-5/National Council for Scientific and Technological Development
- Inova VPPIS-004-FIO-18-52/Fundação Oswaldo Cruz
LinkOut - more resources
Full Text Sources

