SETD8 inhibition targets cancer cells with increased rates of ribosome biogenesis
- PMID: 39341827
- PMCID: PMC11438997
- DOI: 10.1038/s41419-024-07106-6
SETD8 inhibition targets cancer cells with increased rates of ribosome biogenesis
Abstract
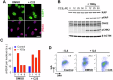
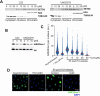
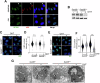
SETD8 is a methyltransferase that is overexpressed in several cancers, which monomethylates H4K20 as well as other non-histone targets such as PCNA or p53. We here report novel SETD8 inhibitors, which were discovered while trying to identify chemicals that prevent 53BP1 foci formation, an event mediated by H4K20 methylation. Consistent with previous reports, SETD8 inhibitors induce p53 expression, although they are equally toxic for p53 proficient or deficient cells. Thermal stability proteomics revealed that the compounds had a particular impact on nucleoli, which was confirmed by fluorescent and electron microscopy. Similarly, Setd8 deletion generated nucleolar stress and impaired ribosome biogenesis, supporting that this was an on-target effect of SETD8 inhibitors. Furthermore, a genome-wide CRISPR screen identified an enrichment of nucleolar factors among those modulating the toxicity of SETD8 inhibitors. Accordingly, the toxicity of SETD8 inhibition correlated with MYC or mTOR activity, key regulators of ribosome biogenesis. Together, our study provides a new class of SETD8 inhibitors and a novel biomarker to identify tumors most likely to respond to this therapy.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

