The chromatin landscape of high-grade serous ovarian cancer metastasis identifies regulatory drivers in post-chemotherapy residual tumour cells
- PMID: 39341888
- PMCID: PMC11438996
- DOI: 10.1038/s42003-024-06909-9
The chromatin landscape of high-grade serous ovarian cancer metastasis identifies regulatory drivers in post-chemotherapy residual tumour cells
Abstract
Disease recurrence following chemotherapy is a major clinical challenge in ovarian cancer (OC), but little is known regarding how the tumour epigenome regulates transcriptional programs underpinning chemoresistance. We determine the single cell chromatin accessibility landscape of omental OC metastasis from treatment-naïve and neoadjuvant chemotherapy-treated patients and define the chromatin accessibility profiles of epithelial, fibroblast, myeloid and lymphoid cells. Epithelial tumour cells display open chromatin regions enriched with motifs for the oncogenic transcription factors MEIS and PBX. Post chemotherapy microenvironments show profound tumour heterogeneity and selection for cells with accessible chromatin enriched for TP53, TP63, TWIST1 and resistance-pathway-activating transcription factor binding motifs. An OC chemoresistant tumour subpopulation known to be present prior to treatment, and characterised by stress-associated gene expression, is enriched post chemotherapy. Nuclear receptors RORa, NR2F6 and HNF4G are uncovered as candidate transcriptional drivers of these cells whilst closure of binding sites for E2F2 and E2F4 indicate post-treated tumour having low proliferative capacity. Delineation of the gene regulatory landscape of ovarian cancer cells surviving chemotherapy treatment therefore reveals potential core transcriptional regulators of chemoresistance, suggesting novel therapeutic targets for improving clinical outcome.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

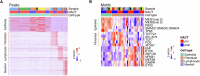




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

