Evolution of aromatic amino acid metabolism in plants: a key driving force behind plant chemical diversity in aromatic natural products
- PMID: 39343022
- PMCID: PMC11439500
- DOI: 10.1098/rstb.2023.0352
Evolution of aromatic amino acid metabolism in plants: a key driving force behind plant chemical diversity in aromatic natural products
Abstract
A diverse array of plant aromatic compounds contributes to the tremendous chemical diversity in the plant kingdom that cannot be seen in microbes or animals. Such chemodiversity of aromatic natural products has emerged, occasionally in a lineage-specific manner, to adopt to challenging environmental niches, as various aromatic specialized metabolites play indispensable roles in plant development and stress responses (e.g. lignin, phytohormones, pigments and defence compounds). These aromatic natural products are synthesized from aromatic amino acids (AAAs), l-tyrosine, l-phenylalanine and l-tryptophan. While amino acid metabolism is generally assumed to be conserved between animals, microbes and plants, recent phylogenomic, biochemical and metabolomic studies have revealed the diversity of the AAA metabolism that supports efficient carbon allocation to downstream biosynthetic pathways of AAA-derived metabolites in plants. This review showcases the intra- and inter-kingdom diversification and origin of committed enzymes involved in plant AAA biosynthesis and catabolism and their potential application as genetic tools for plant metabolic engineering. I also discuss evolutionary trends in the diversification of plant AAA metabolism that expands the chemical diversity of AAA-derived aromatic natural products in plants. This article is part of the theme issue 'The evolution of plant metabolism'.
Keywords: aromatic amino acid; aromatic natural products; enzyme evolution; negative feedback inhibition.
Conflict of interest statement
I declare I have no competing interests.
Figures
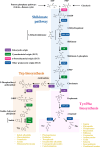
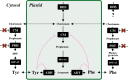
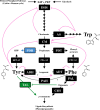
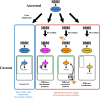
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
