Blending space and time to talk about cancer in extended reality
- PMID: 39343807
- PMCID: PMC11439928
- DOI: 10.1038/s41746-024-01262-x
Blending space and time to talk about cancer in extended reality
Abstract
We introduce a proof-of-concept extended reality (XR) environment for discussing cancer, presenting genomic information from multiple tumour sites in the context of 3D tumour models generated from CT scans. This tool enhances multidisciplinary discussions. Clinicians and cancer researchers explored its use in oncology, sharing perspectives on XR's potential for use in molecular tumour boards, clinician-patient communication, and education. XR serves as a universal language, fostering collaborative decision-making in oncology.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

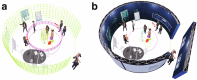
References
Publication types
Grants and funding
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
- 20/765/Manatu Hauora | Health Research Council of New Zealand (HRC)
LinkOut - more resources
Full Text Sources

