Pan-genome analysis of GT64 gene family and expression response to Verticillium wilt in cotton
- PMID: 39343881
- PMCID: PMC11440917
- DOI: 10.1186/s12870-024-05584-6
Pan-genome analysis of GT64 gene family and expression response to Verticillium wilt in cotton
Abstract
Background: The GT64 subfamily, belonging to the glycosyltransferase family, plays a critical function in plant adaptation to stress conditions and the modulation of plant growth, development, and organogenesis processes. However, a comprehensive identification and systematic analysis of GT64 in cotton are still lacking.
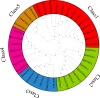
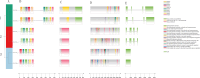
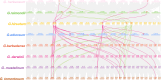
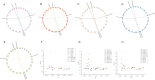
Results: This study used bioinformatics techniques to conduct a detailed investigation on the GT64 gene family members of eight cotton species for the first time. A total of 39 GT64 genes were detected, which could be classified into five subfamilies according to the phylogenetic tree. Among them, six genes were found in upland cotton. Furthermore, investigated the precise chromosomal positions of these genes and visually represented their gene structure details. Moreover, forecasted cis-regulatory elements in GhGT64s and ascertained the duplication type of the GT64 in the eight cotton species. Evaluation of the Ka/Ks ratio for similar gene pairs among the eight cotton species provided insights into the selective pressures acting on these homologous genes. Additionally, analyzed the expression profiles of the GT64 gene family. Overexpressing GhGT64_4 in tobacco improved its disease resistance. Subsequently, VIGS experiments conducted in cotton demonstrated reduced disease resistance upon silencing of the GhGT64_4, may indicate its involvement in affecting lignin and jasmonic acid biosynthesis pathways, thus impacting cotton resistance. Weighted Gene Co-expression Network Analysis (WGCNA) revealed an early immune response against Verticillium dahliae in G. barbadense compared to G. hirsutum. Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR) analysis indicated that some GT64 genes might play a role under various biotic and abiotic stress conditions.
Conclusions: These discoveries enhance our knowledge of GT64 family members and lay the groundwork for future investigations into the disease resistance mechanisms of this gene in cotton.
Keywords: Expression pattern; GT64; Transgenic tobacco; Upland cotton; VIGS; WGCNA.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


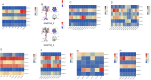
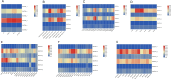


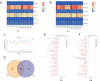


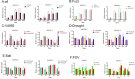
References
MeSH terms
Substances
Grants and funding
- CB2023A19/the National Key Laboratory of Cotton Bio-breeding and Integrated Utilization
- CB2023A19/the National Key Laboratory of Cotton Bio-breeding and Integrated Utilization
- CB2023A19/the National Key Laboratory of Cotton Bio-breeding and Integrated Utilization
- CB2023A19/the National Key Laboratory of Cotton Bio-breeding and Integrated Utilization
- 2019CB008/the Project of Innovation Team Building in Key Areas of Xinjiang Production and Construction Corps
- 2019CB008/the Project of Innovation Team Building in Key Areas of Xinjiang Production and Construction Corps
- 2019CB008/the Project of Innovation Team Building in Key Areas of Xinjiang Production and Construction Corps
- 2019CB008/the Project of Innovation Team Building in Key Areas of Xinjiang Production and Construction Corps
LinkOut - more resources
Full Text Sources

